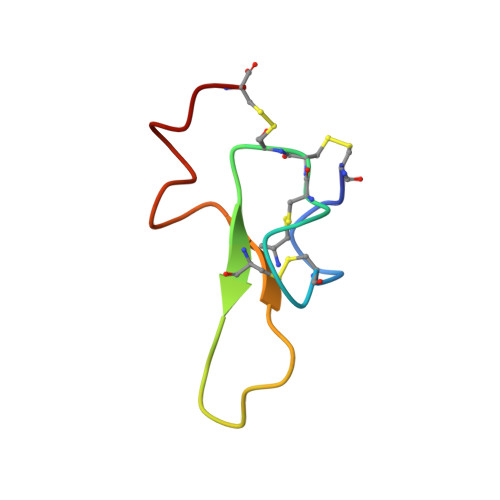
Solution structure of robustoxin, the lethal neurotoxin from the funnel-web spider Atrax robustus.
Pallaghy, P.K., Alewood, D., Alewood, P.F., Norton, R.S.(1997) FEBS Lett 419: 191-196
- PubMed: 9428632 Search on PubMed
- DOI: https://doi.org/10.1016/s0014-5793(97)01452-x
- Primary Citation Related Structures:
1QDP - PubMed Abstract:
The solution structure of robustoxin, the lethal neurotoxin from the Sydney funnel-web spider Atrax robustus, has been determined from 2D 1H NMR data. Robustoxin is a polypeptide of 42 residues cross-linked by four disulphide bonds, the connectivities of which were determined from NMR data and trial structure calculations to be 1-15, 8-20, 14-31 and 16-42 (a 1-4/2-6/3-7/5-8 pattern). The structure consists of a small three-stranded, anti-parallel beta-sheet and a series of interlocking gamma-turns at the C-terminus. It also contains a cystine knot, thus placing it in the inhibitor cystine knot motif family of structures, which includes the omega-conotoxins and a number of plant and animal toxins and protease inhibitors. Robustoxin contains three distinct charged patches on its surface, and an extended loop that includes several aromatic and non-polar residues. Both of these structural features may play a role in its binding to the voltage-gated sodium channel.
- Biomolecular Research Institute, Parkville, Vic., Australia.
Organizational Affiliation: