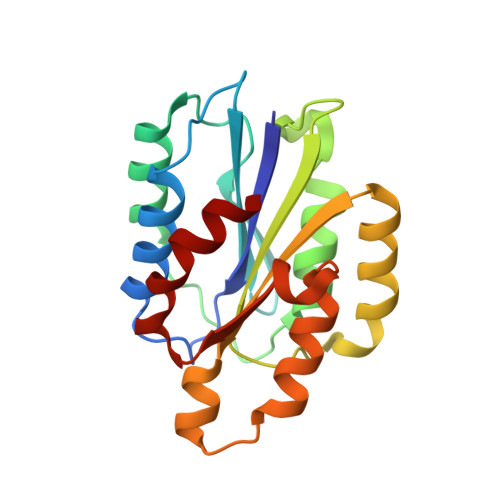
Trench-shaped binding sites promote multiple classes of interactions between collagen and the adherence receptors, alpha(1)beta(1) integrin and Staphylococcus aureus cna MSCRAMM.
Rich, R.L., Deivanayagam, C.C., Owens, R.T., Carson, M., Hook, A., Moore, D., Symersky, J., Yang, V.W., Narayana, S.V., Hook, M.(1999) J Biological Chem 274: 24906-24913
- PubMed: 10455165 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.274.35.24906
- Primary Citation Related Structures:
1QC5 - PubMed Abstract:
Most mammalian cells and some pathogenic bacteria are capable of adhering to collagenous substrates in processes mediated by specific cell surface adherence molecules. Crystal structures of collagen-binding regions of the human integrin alpha(2)beta(1) and a Staphylococcus aureus adhesin reveal a "trench" on the surface of both of these proteins. This trench can accommodate a collagen triple-helical structure and presumably represents the ligand-binding site (Emsley, J., King, S. L., Bergelson, J. M., and Liddington, R. C. (1997) J. Biol. Chem. 272, 28512-28517; Symersky, J., Patti, J. M., Carson, M., House-Pompeo, K., Teale, M., Moore, D., Jin, L., Schneider, A., DeLucas, L. J., Höök, M., and Narayana, S. V. L. (1997) Nat. Struct. Biol. 4, 833-838). We report here the crystal structure of the alpha subunit I domain from the alpha(1)beta(1) integrin. This collagen-binding protein also contains a trench on one face in which the collagen triple helix may be docked. Furthermore, we compare the collagen-binding mechanisms of the human alpha(1) integrin I domain and the A domain from the S. aureus collagen adhesin, Cna. Although the S. aureus and human proteins have unrelated amino acid sequences, secondary structure composition, and cation requirements for effective ligand binding, both proteins bind at multiple sites within one collagen molecule, with the sites in collagen varying in their affinity for the adherence molecule. We propose that (i) these evolutionarily dissimilar adherence proteins recognize collagen via similar mechanisms, (ii) the multisite, multiclass protein/ligand interactions observed in these two systems result from a binding-site trench, and (iii) this unusual binding mechanism may be thematic for proteins binding extended, rigid ligands that contain repeating structural motifs.
- Center for Extracellular Matrix Biology, Institute of Biosciences and Technology, Texas A&M University, Houston, Texas 77030, USA. rrich@ibt.tamu.edu
Organizational Affiliation: