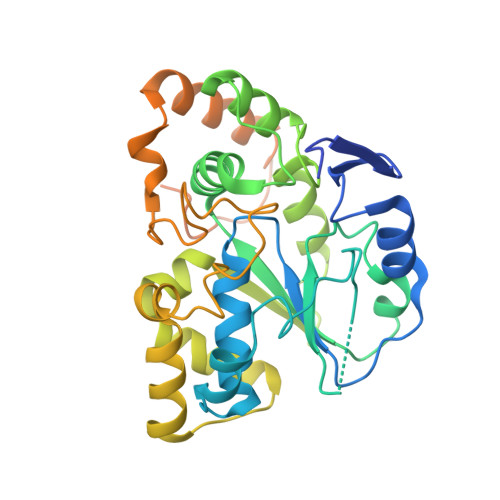
Crystal structure of human cholesterol sulfotransferase (SULT2B1b) in the presence of pregnenolone and 3'-phosphoadenosine 5'-phosphate. Rationale for specificity differences between prototypical SULT2A1 and the SULT2BG1 isoforms.
Lee, K.A., Fuda, H., Lee, Y.C., Negishi, M., Strott, C.A., Pedersen, L.C.(2003) J Biological Chem 278: 44593-44599
- PubMed: 12923182 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M308312200
- Primary Citation Related Structures:
1Q1Q, 1Q1Z, 1Q20, 1Q22 - PubMed Abstract:
The gene for human hydroxysteroid sulfotransferase (SULT2B1) encodes two peptides, SULT2B1a and SULT2B1b, that differ only at their amino termini. SULT2B1b has a predilection for cholesterol but is also capable of sulfonating pregnenolone, whereas SULT2B1a preferentially sulfonates pregnenolone and only minimally sulfonates cholesterol. We have determined the crystal structure of SULT2B1a and SULT2B1b bound to the substrate donor product 3'-phosphoadenosine 5'-phosphate at 2.9 and 2.4 A, respectively, as well as SULT2B1b in the presence of the acceptor substrate pregnenolone at 2.3 A. These structures reveal a different catalytic binding orientation for the substrate from a previously determined structure of hydroxysteroid sulfotransferase (SULT2A1) binding dehydroepiandrosterone. In addition, the amino-terminal helix comprising residues Asp19 to Lys26, which determines the specificity difference between the SULT2B1 isoforms, becomes ordered upon pregnenolone binding, covering the substrate binding pocket.
- Laboratory of Structural Biology, National Institute of Environmental Health Sciences, National Institutes of Health, Research Triangle Park, North Carolina 27709, USA.
Organizational Affiliation: