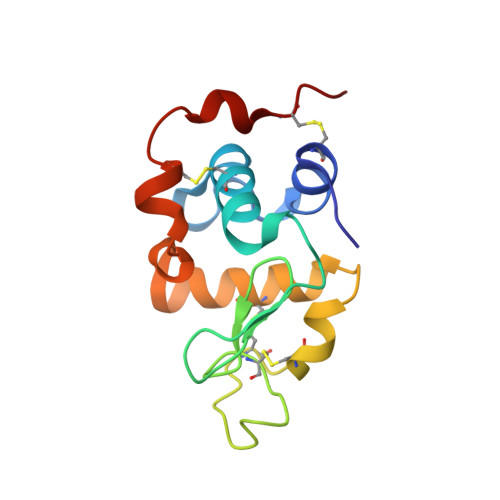
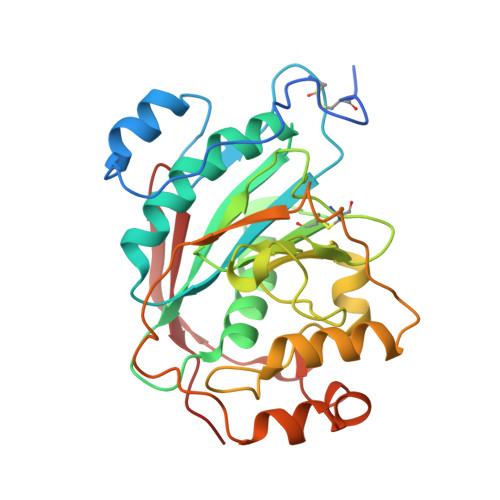
The Role of Tryptophan 314 in the Conformational Changes of beta 1,4-Galactosyltransferase-I
Ramasamy, V., Ramakrishnan, B., Boeggeman, E., Qasba, P.K.(2003) J Mol Biology 331: 1065-1076
- PubMed: 12927542 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(03)00790-3
- Primary Citation Related Structures:
1PZT, 1PZY - PubMed Abstract:
beta1,4-Galactosyltransferase-I (beta4Gal-T1) undergoes critical conformational changes upon substrate binding from an open conformation (conf-I) to the closed conformation (conf-II). This change involves two flexible loops: the small (residues 313-316) and the long loop (residues 345-365). Upon substrate binding, Trp314 in the small flexible loop moves towards the catalytic pocket and interacts with the donor and the acceptor substrates. For a better understanding of the role played by Trp314 in the conformational changes of beta4Gal-T1, we mutated it to Ala and carried out substrate-binding, proteolytic and crystallographic studies. The W314A mutation reduces the enzymatic activity, binding to substrates and to the modifier protein, alpha-lactalbumin (LA), by over 99%. The limited proteolysis with Glu-C or Lys-C proteases shows differences in the rate of cleavage of the long loop of the wild-type and mutant W314A, indicating conformational differences in the region between the two proteins. Without substrate, the mutant crystallizes in a conformation (conf-I') (1.9A resolution crystal structure), that is not identical with, but close to an open conformation (conf-I), whereas its complex with the substrates and alpha-lactalbumin, crystallizes in a conformation (2.3A resolution crystal structure) that is identical with the closed conformation (conf-II). This study shows the crucial role Trp314 plays in the conformational state of the long loop, in the binding of substrates and in the catalytic mechanism of the enzyme.
- Structural Glycobiology Section, LECB, CCR, NCI-Frederick, Building 469, Room 221, 21702, Frederick, MD, USA.
Organizational Affiliation: