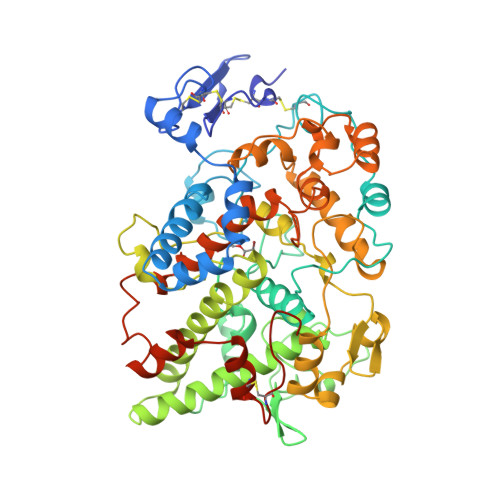
The structural basis of aspirin activity inferred from the crystal structure of inactivated prostaglandin H2 synthase.
Loll, P.J., Picot, D., Garavito, R.M.(1995) Nat Struct Biol 2: 637-643
- PubMed: 7552725
- DOI: https://doi.org/10.1038/nsb0895-637
- Primary Citation Related Structures:
1PTH - PubMed Abstract:
Aspirin exerts its anti-inflammatory effects through selective acetylation of serine 530 on prostaglandin H2 synthase (PGHS). Here we present the 3.4 A resolution X-ray crystal structure of PGHS isoform-1 inactivated by the potent aspirin analogue 2-bromoacetoxy-benzoic acid. Acetylation by this analogue abolishes cyclooxygenase activity by steric blockage of the active-site channel and not through a large conformational change. We observe two rotameric states of the acetyl-serine side chain which block the channel to different extents, a result which may explain the dissimilar effects of aspirin on the two PGHS isoforms. We also observe the product salicylic acid binding at a site consistent with its antagonistic effect on aspirin activity.
- Department of Biochemistry and Molecular Biology, University of Chicago, Illinois 60637, USA.
Organizational Affiliation: