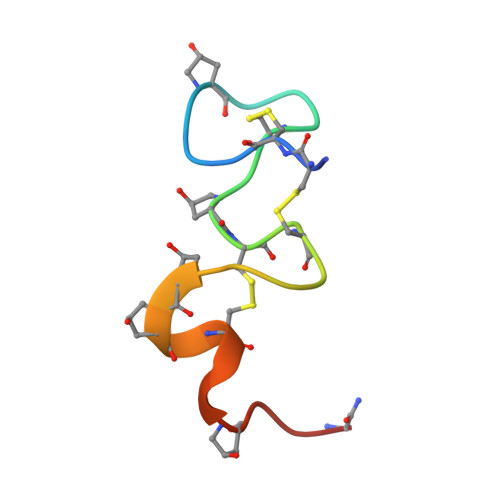
Solution Conformation of alphaA-conotoxin EIVA, a Potent Neuromuscular Nicotinic Acetylcholine Receptor Antagonist from Conus ermineus
Chi, S.-W., Park, K.-H., Suk, J.-E., Olivera, B.M., McIntosh, J.M., Han, K.-H.(2003) J Biological Chem 278: 42208-42213
- PubMed: 12900418
- DOI: https://doi.org/10.1074/jbc.M303342200
- Primary Citation of Related Structures:
1PQR - PubMed Abstract:
We report the solution three-dimensional structure of an alphaA-conotoxin EIVA determined by nuclear magnetic resonance spectroscopy and restrained molecular dynamics. The alphaA-conotoxin EIVA consists of 30 amino acids representing the largest peptide among the alpha/alphaA-family conotoxins discovered so far and targets the neuromuscular nicotinic acetylcholine receptor with high affinity. alphaA-Conotoxin EIVA consists of three distinct structural domains. The first domain is mainly composed of the Cys3-Cys11-disulfide loop and is structurally ill-defined with a large backbone root mean square deviation of 1.91 A. The second domain formed by residues His12-Hyp21 is extremely well defined with a backbone root mean square deviation of 0.52 A, thus forming a sturdy stem for the entire molecule. The third C-terminal domain formed by residues Hyp22-Gly29 shows an intermediate structural order having a backbone root mean square deviation of 1.04 A. A structurally ill-defined N-terminal first loop domain connected to a rigid central molecular stem seems to be the general structural feature of the alphaA-conotoxin subfamily. A detailed structural comparison between alphaA-conotoxin EIVA and alphaA-conotoxin PIVA suggests that the higher receptor affinity of alphaA-conotoxin EIVA than alphaA-conotoxin PIVA might originate from different steric disposition and charge distribution in the second loop "handle" motif.
- Department of Biology, University of Utah, Salt Lake City, Utah 84112, USA.
Organizational Affiliation: