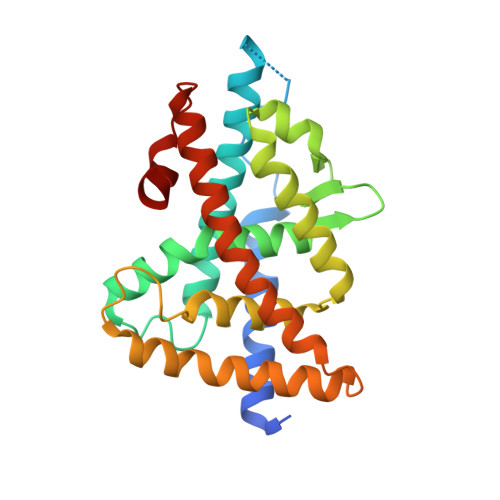
The three-dimensional structure of the liver X receptor beta reveals a flexible ligand-binding pocket that can accommodate fundamentally different ligands.
Farnegardh, M., Bonn, T., Sun, S., Ljunggren, J., Ahola, H., Wilhelmsson, A., Gustafsson, J.-A., Carlquist, M.(2003) J Biological Chem 278: 38821-38828
- PubMed: 12819202 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M304842200
- Primary Citation Related Structures:
1PQ6, 1PQ9, 1PQC - PubMed Abstract:
The structures of the liver X receptor LXRbeta (NR1H2) have been determined in complexes with two synthetic ligands, T0901317 and GW3965, to 2.1 and 2.4 A, respectively. Together with its isoform LXRalpha (NR1H3) it regulates target genes involved in metabolism and transport of cholesterol and fatty acids. The two LXRbeta structures reveal a flexible ligand-binding pocket that can adjust to accommodate fundamentally different ligands. The ligand-binding pocket is hydrophobic but with polar or charged residues at the two ends of the cavity. T0901317 takes advantage of this by binding to His-435 close to H12 while GW3965 orients itself with its charged group in the opposite direction. Both ligands induce a fixed "agonist conformation" of helix H12 (also called the AF-2 domain), resulting in a transcriptionally active receptor.
- Karo Bio AB, NOVUM, Karolinska Institute, Huddinge University Hosptial, Sweden. mathias.farnegardh@karobio.se
Organizational Affiliation: