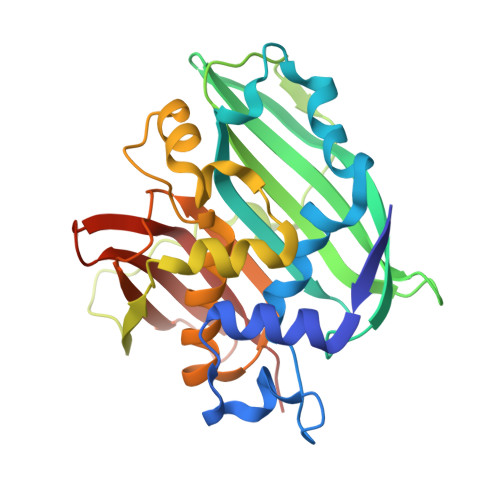
A Two-domain Structure of One Subunit Explains Unique Features of Eukaryotic Hydratase 2.
Koski, M.K., Haapalainen, A.M., Hiltunen, J.K., Glumoff, T.(2004) J Biological Chem 279: 24666-24672
- PubMed: 15051722
- DOI: https://doi.org/10.1074/jbc.M400293200
- Primary Citation Related Structures:
1PN2, 1PN4 - PubMed Abstract:
2-Enoyl-CoA hydratase 2, a part from multifunctional enzyme type 2, hydrates trans-2-enoyl-CoA to 3-hydroxyacyl-CoA in the (3R)-hydroxy-dependent route of peroxisomal beta-oxidation of fatty acids. Unliganded and (3R)-hydroxydecanoyl coenzyme A-complexed crystal structures of 2-enoyl-CoA hydratase 2 from Candida tropicalis multifunctional enzyme type 2 were solved to 1.95- and 2.35-A resolution, respectively. 2-Enoyl-CoA hydratase 2 is a dimeric, alpha+beta protein with a novel quaternary structure. The overall structure of the two-domain subunit of eukaryotic 2-enoyl-CoA hydratase 2 resembles the homodimeric, hot dog fold structures of prokaryotic (R)-specific 2-enoyl-CoA hydratase and beta-hydroxydecanoyl thiol ester dehydrase. Importantly, though, the eukaryotic hydratase 2 has a complete hot dog fold only in its C-domain, whereas the N-domain lacks a long central alpha-helix, thus creating space for bulkier substrates in the binding pocket and explaining the observed difference in substrate preference between eukaryotic and prokaryotic enzymes. Although the N- and C-domains have an identity of <10% at the amino acid level, they share a 50% identity at the nucleotide level and fold similarly. We suggest that a subunit of 2-enoyl-CoA hydratase 2 has evolved via a gene duplication with the concomitant loss of one catalytic site. The hydrogen bonding network of the active site of 2-enoyl-CoA hydratase 2 resembles the active site geometry of mitochondrial (S)-specific 2-enoyl-CoA hydratase 1, although in a mirror image fashion. This arrangement allows the reaction to occur by similar mechanism, supported by mutagenesis and mechanistic studies, although via reciprocal stereochemistry.
- Department of Biochemistry and Biocenter Oulu, University of Oulu, P. O. Box 3000, FIN-90014 University of Oulu, Finland.
Organizational Affiliation: