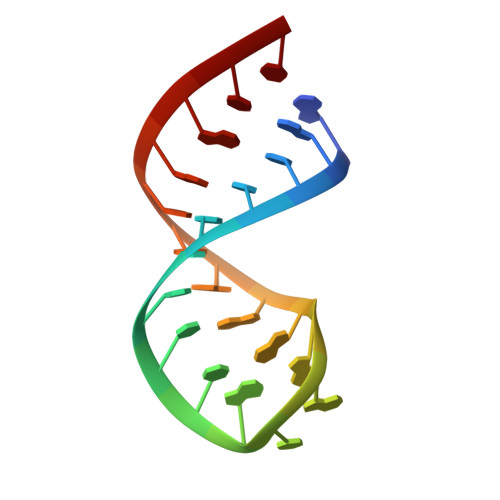
Solution structure of the HIV-1 frameshift inducing stem-loop RNA.
Staple, D.W., Butcher, S.E.(2003) Nucleic Acids Res 31: 4326-4331
- PubMed: 12888491
- DOI: https://doi.org/10.1093/nar/gkg654
- Primary Citation of Related Structures:
1PJY - PubMed Abstract:
The translation of reverse transcriptase and other essential viral proteins from the HIV-1 Pol mRNA requires a programmed -1 ribosomal frameshift. This frameshift is induced by two highly conserved elements within the HIV-1 mRNA: a slippery sequence comprised of a UUUUUUA heptamer, and a downstream stem-loop structure. We have determined the structure of the HIV-1 frameshift inducing RNA stem-loop, using multidimensional heteronuclear nuclear magnetic resonance (NMR) methods. The 22 nucleotide RNA solution structure [root mean squared deviation (r.m.s.d.) = 1.2 A] was determined from 475 nuclear Overhauser effect (NOE)-derived distance restrains, 20 residual dipolar couplings and direct detection of hydrogen bonds via scalar couplings. We find that the frameshift inducing stem-loop is an A-form helix capped by a structured ACAA tetraloop. The ACAA tetraloop is stabilized by an equilateral 5' and 3' stacking pattern, a sheared A-A pair and a cross-strand hydrogen bond. Unexpectedly, the ACAA tetraloop structure is nearly identical to a known tetraloop fold, previously identified in the RNase III recognition site from Saccharomyces cerevisiae.
- Department of Biochemistry, University of Wisconsin-Madison, 433 Babcock Drive, Madison, WI 53706, USA.
Organizational Affiliation: