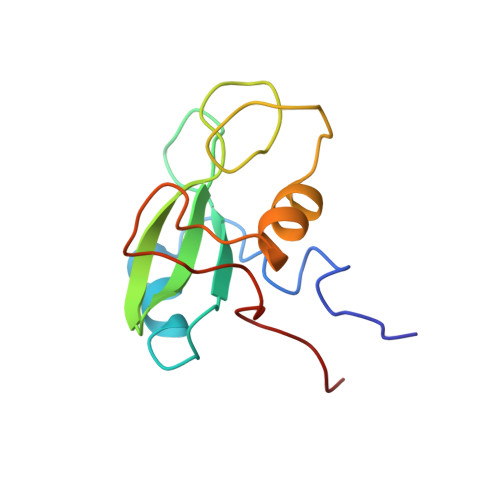
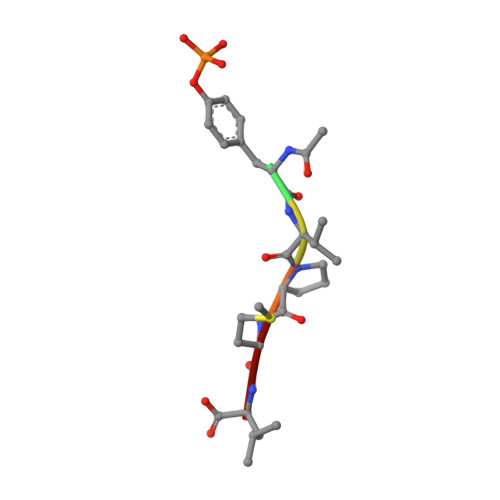
Structure of a specific peptide complex of the carboxy-terminal SH2 domain from the p85 alpha subunit of phosphatidylinositol 3-kinase.
Breeze, A.L., Kara, B.V., Barratt, D.G., Anderson, M., Smith, J.C., Luke, R.W., Best, J.R., Cartlidge, S.A.(1996) EMBO J 15: 3579-3589
- PubMed: 8670861 Search on PubMedSearch on PubMed Central
- Primary Citation Related Structures:
1PIC - PubMed Abstract:
We have determined the solution structure of the C-terminal SH2 domain of the p85 alpha subunit of human phosphatidylinositol (PI) 3-kinase (EC 2.7.1.137) in complex with a phosphorylated tyrosine pentapeptide sequence from the platelet-derived growth factor receptor using heteronuclear nuclear magnetic resonance spectroscopy. Overall, the structure is similar to other SH2 domain complexes, but displays different detail interactions within the phosphotyrosine binding site and in the recognition site for the +3 methionine residue of the peptide, the side chain of which inserts into a particularly deep and narrow pocket which is displaced relative to that of other SH2 domains. The contacts made within this +3 pocket provide the structural basis for the strong selection for methionine at this position which characterizes the SH2 domains of PI3-kinase. Comparison with spectral and structural features of the uncomplexed domain shows that the long BG loop becomes less mobile in the presence of the bound peptide. In contrast, extreme resonance broadening encountered for most residues in the beta D', beta E and beta F strands and associated connecting loops of the domain in the absence of peptide persists in the complex, implying conformational averaging in this part of the molecule on a microsecond-to-millisecond time scale.
- Protein Structure Laboratory, Zeneca Pharmaceuticals, Mereside, Alderley Park, Cheshire, UK.
Organizational Affiliation: