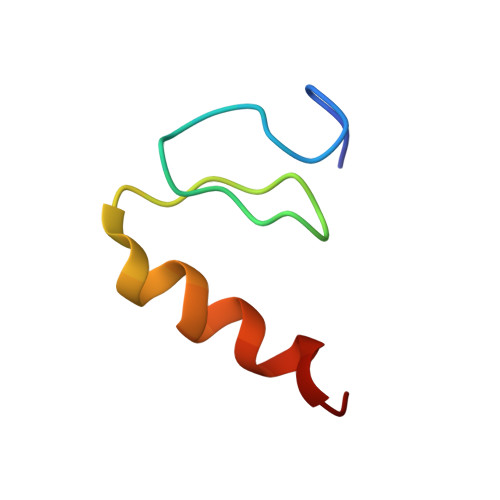
CCHX zinc finger derivatives retain the ability to bind Zn(II) and mediate protein-DNA interactions.
Simpson, R.J., Cram, E.D., Czolij, R., Matthews, J.M., Crossley, M., Mackay, J.P.(2003) J Biological Chem 278: 28011-28018
- PubMed: 12736264 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M211146200
- Primary Citation Related Structures:
1P7A - PubMed Abstract:
Classical (CCHH) zinc fingers are among the most common protein domains found in eukaryotes. They function as molecular recognition elements that mediate specific contact with DNA, RNA, or other proteins and are composed of a betabetaalpha fold surrounding a single zinc ion that is ligated by two cysteine and two histidine residues. In a number of variant zinc fingers, the final histidine is not conserved, and in other unrelated zinc binding domains, residues such as aspartate can function as zinc ligands. To test whether the final histidine is required for normal folding and the DNA-binding function of classical zinc fingers, we focused on finger 3 of basic Krüppel-like factor. The structure of this domain was determined using NMR spectroscopy and found to constitute a typical classical zinc finger. We generated a panel of substitution mutants at the final histidine in this finger and found that several of the mutants retained some ability to fold in the presence of zinc. Consistent with this result, we showed that mutation of the final histidine had only a modest effect on DNA binding in the context of the full three-finger DNA-binding domain of basic Krüppel-like factor. Further, the zinc binding ability of one of the point mutants was tested and found to be indistinguishable from the wild-type domain. These results suggest that the final zinc chelating histidine is not an essential feature of classical zinc fingers and have implications for zinc finger evolution, regulation, and the design of experiments testing the functional roles of these domains.
- School of Molecular and Microbial Biosciences, G08, University of Sydney, New South Wales 2006, Australia.
Organizational Affiliation: