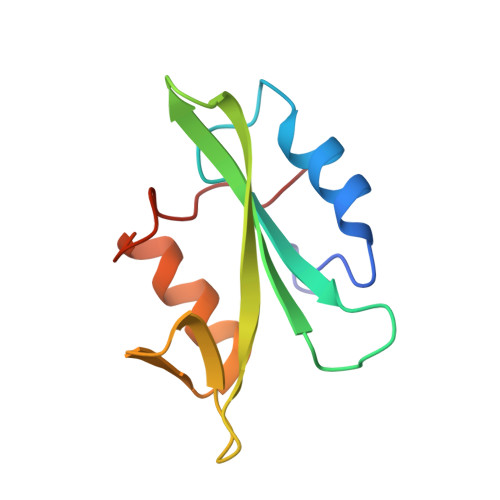
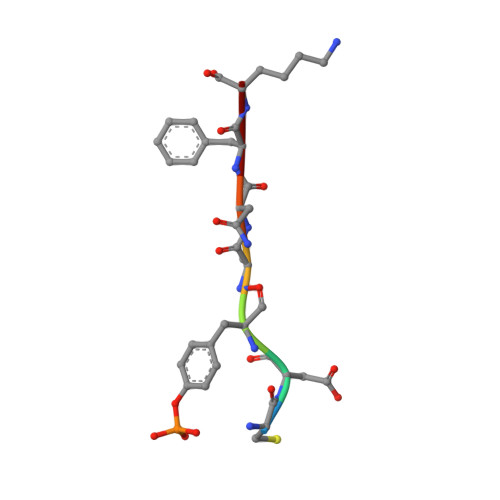
The structure of the membrane distal phosphatase domain of RPTPalpha reveals interdomain flexibility and an SH2 domain interaction region.
Sonnenburg, E.D., Bilwes, A., Hunter, T., Noel, J.P.(2003) Biochemistry 42: 7904-7914
- PubMed: 12834342 Search on PubMed
- DOI: https://doi.org/10.1021/bi0340503
- Primary Citation Related Structures:
1P13, 1P15 - PubMed Abstract:
The receptor protein tyrosine phosphatase alpha (RPTPalpha) is a transmembrane receptor with two intracellular protein tyrosine phosphatase domains, a catalytically active membrane proximal domain (D1) and a membrane distal phosphatase domain with minimal catalytic activity (D2). Here we elucidate the crystal structure of RPTPalpha's D2 domain. Unlike D1, D2 exists as a monomer and lacks the N-terminal inhibitory wedge motif. The N-terminal portion of D2 is disordered, and this region linking D1 to D2 is proteolytically labile in solution whether part of D2 alone or tethered to D1, indicating that the polypeptide backbone of this part of D2 is highly flexible, and therefore accessible to proteases under native conditions. Furthermore, we have crystallized the SH2 domain of the protein tyrosine kinase c-Src, a RPTPalpha substrate, with a phosphopeptide encompassing the C-terminal phosphorylation site of D2 (pTyr789). The SH2 domain of Src binds RPTPalpha in an extended conformation. The structural and functional data support a D1-D2 arrangement with significant flexibility between phosphatase domains of RPTPalpha that is likely to be important for dynamic alterations in intra- and/or intermolecular interactions that are critical for RPTPalpha function.
- Structural Biology Laboratory, The Salk Institute for Biological Studies, 10010 North Torrey Pines Road, La Jolla, California 92037, USA.
Organizational Affiliation: