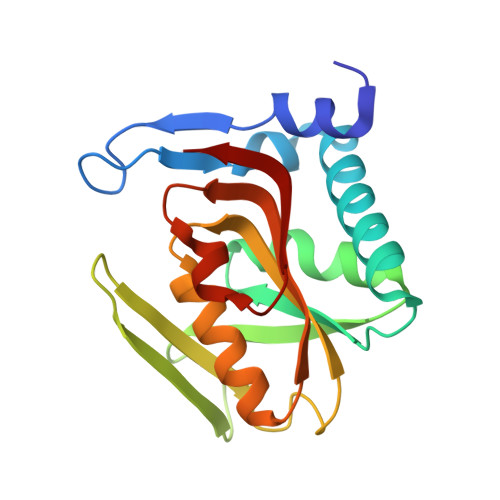
Three-dimensional structure of human adenine phosphoribosyltransferase and its relation to DHA-urolithiasis.
Silva, M., Silva, C.H.T.P., Iulek, J., Thiemann, O.H.(2004) Biochemistry 43: 7663-7671
- PubMed: 15196008 Search on PubMed
- DOI: https://doi.org/10.1021/bi0360758
- Primary Citation Related Structures:
1ORE - PubMed Abstract:
In mammals, adenine phosphoribosyltransferase (APRT, EC 2.4.2.7) is present in all tissues and provides the only known mechanism for the metabolic salvage of adenine resulting from the polyamine biosynthesis pathway or from dietary sources. In humans, APRT deficiency results in serious kidney illness such as nephrolithiasis, interstitial nephritis, and chronic renal failure as a result of 2,8-dihydroxyadenine (DHA) precipitation in the renal interstitium. To address the molecular basis of DHA-urolithiasis, the recombinant human APRT was crystallized in complex with adenosine 5'-monophosphate (AMP). Refinement of X-ray diffraction data extended to 2.1 A resolution led to a final crystallographic R(factor) of 13.3% and an R(free) of 17.6%. This structure is composed of nine beta-strands and six alpha-helices, and the active site pocket opens slightly to accommodate the AMP product. The core of APRT is similar to that of other phosphoribosyltransferases (PRTases), although the adenine-binding domain is quite different. Structural comparisons between the human APRT and other "type I" PRTases of known structure revealed several important features of the biochemistry of PRTases. We propose that the residues located at positions corresponding to Leu159 and Ala131 in hAPRT are responsible for the base specificities of type I PRTases. The comparative analysis shown here also provides structural information for the mechanism by which mutations in the human APRT lead to DHA-urolithiasis.
- Laboratory of Protein Crystallography and Structural Biology, Physics Institute of São Carlos, University of São Paulo, Av. Trabalhador Sãocarlense 400, P.O. Box 369, 13566-590 São Carlos, SP, Brazil.
Organizational Affiliation: