Modulation of alpha-thrombin function by distinct interactions with platelet glycoprotein Ibalpha
Celikel, R., McClintock, R.A., Roberts, J.R., Mendolicchio, G.L., Ware, J., Varughese, K.I., Ruggeri, Z.M.(2003) Science 301: 218-221
- PubMed: 12855810 Search on PubMed
- DOI: https://doi.org/10.1126/science.1084183
- Primary Citation Related Structures:
1OOK, 1P9A - PubMed Abstract:
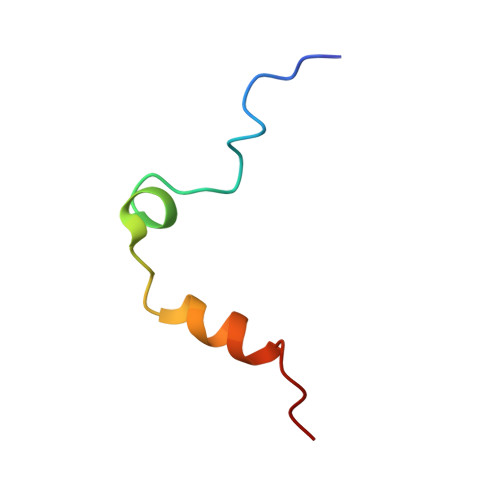
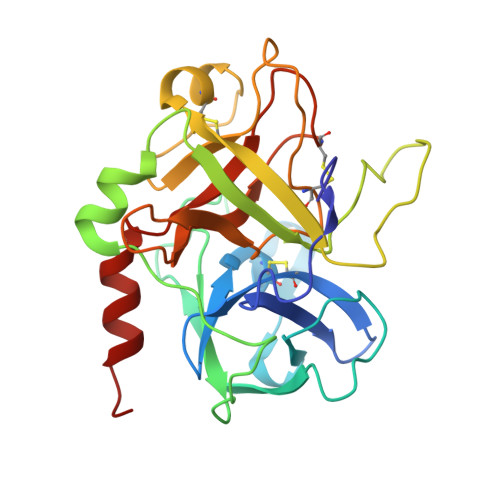
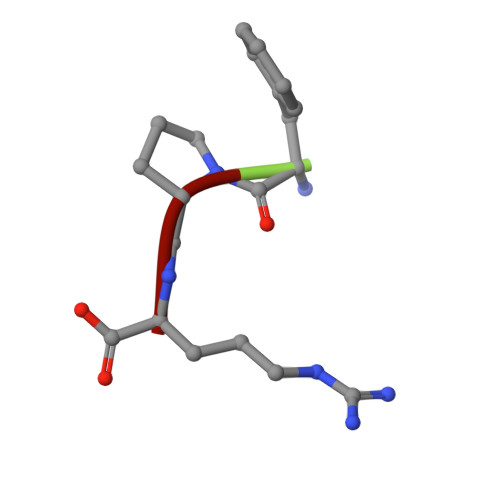
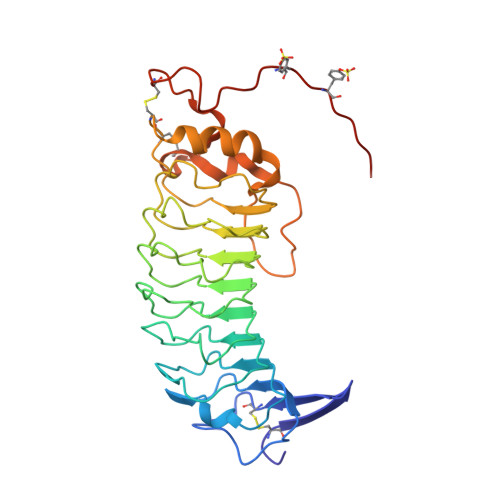
Thrombin bound to platelets contributes to stop bleeding and, in pathological conditions, may cause vascular thrombosis. We have determined the structure of platelet glycoprotein Ibalpha (GpIbalpha) bound to thrombin at 2.3 angstrom resolution and defined two sites in GpIbalpha that bind to exosite II and exosite I of two distinct alpha-thrombin molecules, respectively. GpIbalpha occupancy may be sequential, as the site binding to alpha-thrombin exosite I appears to be cryptic in the unoccupied receptor but exposed when a first thrombin molecule is bound through exosite II. These interactions may modulate alpha-thrombin function by mediating GpIbalpha clustering and cleavage of protease-activated receptors, which promote platelet activation, while limiting fibrinogen clotting through blockade of exosite I.
- Roon Research Center for Arteriosclerosis and Thrombosis, Division of Experimental Thrombosis and Hemostasis, Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: