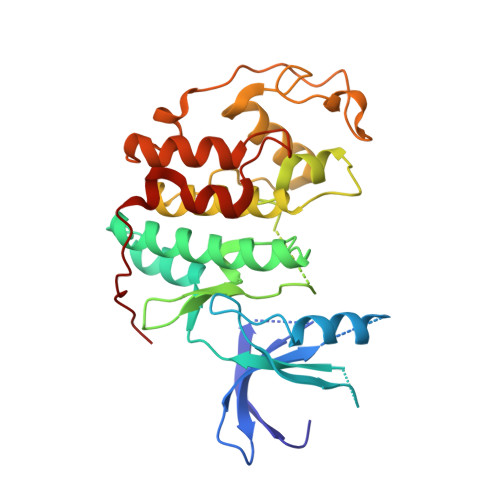
Imidazo[1,2-A]Pyridines: A Potent and Selective Class of Cyclin-Dependent Kinase Inhibitors Identified Through Structure-Based Hybridisation
Anderson, M., Beattie, J.F., Breault, G.A., Breed, J., Byth, K.F., Culshaw, J.D., Ellston, R.P.A., Green, S., Minshull, C.A., Norman, R.A., Pauptit, R.A., Stanway, J., Thomas, A.P., Jewsbury, P.J.(2003) Bioorg Med Chem Lett 13: 3021
- PubMed: 12941325 Search on PubMed
- DOI: https://doi.org/10.1016/s0960-894x(03)00638-3
- Primary Citation Related Structures:
1OIQ, 1OIR, 1OIT - PubMed Abstract:
High-throughput screening identified the imidazo[1,2-a]pyridine and bisanilinopyrimidine series as inhibitors of the cyclin-dependent kinase CDK4. Comparison of their experimentally-determined binding modes and emerging structure-activity trends led to the development of potent and selective imidazo[1,2-a]pyridine inhibitors for CDK4 and in particular CDK2.
- AstraZeneca, Alderley Park, Macclesfield, Cheshire SK10 4TG, UK.
Organizational Affiliation: