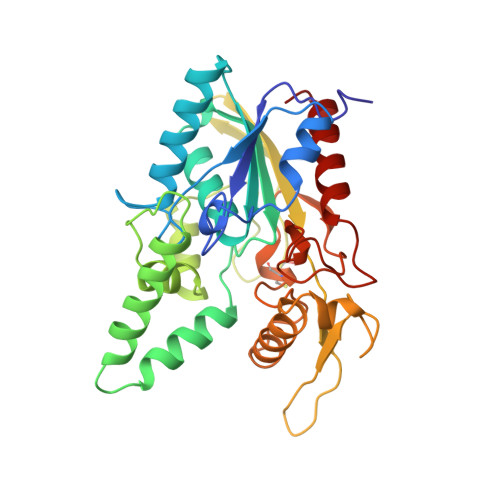
The crystal structure of a triacylglycerol lipase from Pseudomonas cepacia reveals a highly open conformation in the absence of a bound inhibitor.
Kim, K.K., Song, H.K., Shin, D.H., Hwang, K.Y., Suh, S.W.(1997) Structure 5: 173-185
- PubMed: 9032073 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(97)00177-9
- Primary Citation Related Structures:
1OIL - PubMed Abstract:
. Lipases, a family of enzymes which catalyze the hydrolysis of triglycerides, are widely distributed in many organisms. True lipases are distinguished from esterases by the characteristic interfacial activation they exhibit at an oil-water interface. Lipases are one of the most frequently used biocatalysts for organic reactions performed under mild conditions. Their biotechnological applications include food and oil processing and the preparation of chiral intermediates for the synthesis of enantiomerically pure pharmaceuticals. Recent structural studies on several lipases have provided some clues towards understanding the mechanisms of hydrolytic activity, interfacial activation, and stereoselectivity. This study was undertaken in order to provide structural information on bacterial lipases, which is relatively limited in comparison to that on the enzymes from other sources. . We have determined the crystal structure of a triacylglycerol lipase from Pseudomonas cepacia (PcL) in the absence of a bound inhibitor using X-ray crystallography. The structure shows the lipase to contain an alpha/beta-hydrolase fold and a catalytic triad comprising of residues Ser87, His286 and Asp264. The enzyme shares several structural features with homologous lipases from Pseudomonas glumae (PgL) and Chromobacterium viscosum (CvL), including a calcium-binding site. The present structure of PcL reveals a highly open conformation with a solvent-accessible active site. This is in contrast to the structures of PgL and PcL in which the active site is buried under a closed or partially opened 'lid', respectively. . PcL exhibits some structural features found in other lipases. The presence of the Ser-His-Asp catalytic triad, an oxyanion hole, and the opening of a helical lid suggest that this enzyme shares the same mechanisms of catalysis and interfacial activation as other lipases. The highly open conformation observed in this study is likely to reflect the activated form of the lipase at an oil-water interface. The structure suggests that the interfacial activation of bacterial lipases involves the reorganization of secondary structures and a large movement of the lid to expose the active site. This is similar to the mechanism described for other well characterized fungal and mammalian lipases.
- Department of Chemistry, College of Natural Sciences, Seoul National University, Seoul 151-742, Korea.
Organizational Affiliation: