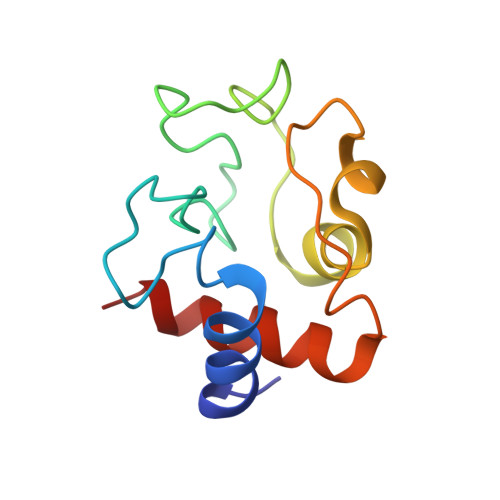
Solution structure of horse heart ferricytochrome c and detection of redox-related structural changes by high-resolution 1H NMR.
Qi, P.X., Beckman, R.A., Wand, A.J.(1996) Biochemistry 35: 12275-12286
- PubMed: 8823161 Search on PubMed
- DOI: https://doi.org/10.1021/bi961042w
- Primary Citation Related Structures:
1OCD, 2FRC - PubMed Abstract:
A model for the solution structure of horse heart ferricytochrome c has been determined by nuclear magnetic resonance spectroscopy combined with hybrid distance geometry-simulated annealing calculations. Forty-four highly refined structures were obtained using a total of 1671 distance constraints based on the observed magnitude of nuclear Overhauser effects and 58 torsion angle restrains based on the magnitude of determined J-coupling constants. The model incorporates six long-lived water molecules detected by pseudo-two-dimensional NOESY-TOCSY spectra. The all-residue root mean square deviation about the average structure is 0.33 +/- 0.04 A for the backbone N, C alpha, and C' atoms and 0.83 +/- 0.05 A for all heavy atoms. The overall topology of the model for solution structure is very similar to that seen in previously reported models for crystal structures of homologous c-type cytochromes though there are a number of significant differences in detailed aspects of the structure. Two of the three main helices display localized irregularities in helical hydrogen bonding resulting in bifurcation of main chain hydrogen bond acceptor carbonyls. The N- and C-terminal helices are tightly packed and display several interhelical interactions not seen in reported crystal models. To provide an independent measure of the accuracy of the model for the oxidized protein, the expected pseudocontact shifts induced by the spin 1/2 iron were compared to the observed redox-dependent chemical shift changes. These comparisons confirm the general accuracy of the model for the oxidized protein and its observed differences with the structure of the reduced protein. The structures of the reduced and oxidized states of the protein provide a template to explain a range of physical and biological data spanning the redox properties, folding, molecular recognition, and stability of the cytochrome c molecule. For example, a redox-dependent reorganization of surface residues at the heme edge can be directly related to the redox behavior of the protein and thereby provides a previously undocumented linkage between structural change potentially associated with molecular recognition of redox partners and the fundamental parameters governing electron transfer.
- Department of Biological Sciences, State University of New York at Buffalo 14260-3000, USA.
Organizational Affiliation: