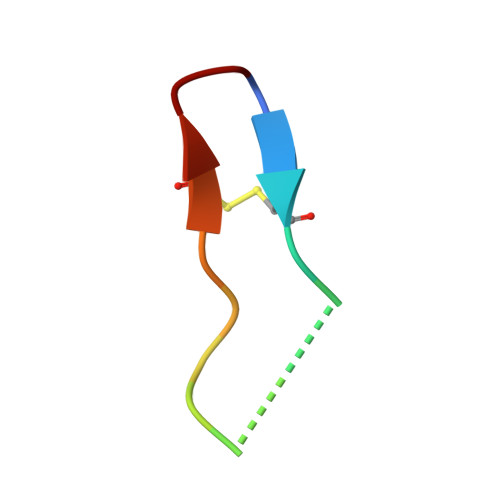
Enzymatic Cyclization of a Potent Bowman-Birk Protease Inhibitor, Sunflower Trypsin Inhibitor-1, and Solution Structure of an Acyclic Precursor Peptide
Marx, U.C., Korsinczky, M., Schirra, H., Jones, A., Condie, B., Otvos, L., Craik, D.J.(2003) J Biological Chem 278: 21782
- PubMed: 12621047 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M212996200
- Primary Citation Related Structures:
1O8Y, 1O8Z - PubMed Abstract:
The most potent known naturally occurring Bowman-Birk inhibitor, sunflower trypsin inhibitor-1 (SFTI-1), is a bicyclic 14-amino acid peptide from sunflower seeds comprising one disulfide bond and a cyclic backbone. At present, little is known about the cyclization mechanism of SFTI-1. We show here that an acyclic permutant of SFTI-1 open at its scissile bond, SFTI-1[6,5], also functions as an inhibitor of trypsin and that it can be enzymatically backbone-cyclized by incubation with bovine beta-trypsin. The resulting ratio of cyclic SFTI-1 to SFTI-1[6,5] is approximately 9:1 regardless of whether trypsin is incubated with SFTI-1[6,5] or SFTI-1. Enzymatic resynthesis of the scissile bond to form cyclic SFTI-1 is a novel mechanism of cyclization of SFTI-1[6,5]. Such a reaction could potentially occur on a trypsin affinity column as used in the original isolation procedure of SFTI-1. We therefore extracted SFTI-1 from sunflower seeds without a trypsin purification step and confirmed that the backbone of SFTI-1 is indeed naturally cyclic. Structural studies on SFTI-1[6,5] revealed high heterogeneity, and multiple species of SFTI-1[6,5] were identified. The main species closely resembles the structure of cyclic SFTI-1 with the broken binding loop able to rotate between a cis/trans geometry of the I7-P8 bond with the cis conformer being similar to the canonical binding loop conformation. The non-reactive loop adopts a beta-hairpin structure as in cyclic wild-type SFTI-1. Another species exhibits an iso-aspartate residue at position 14 and provides implications for possible in vivo cyclization mechanisms.
- Institute for Molecular Bioscience, University of Queensland, Brisbane, Queensland 4072, Australia.
Organizational Affiliation: