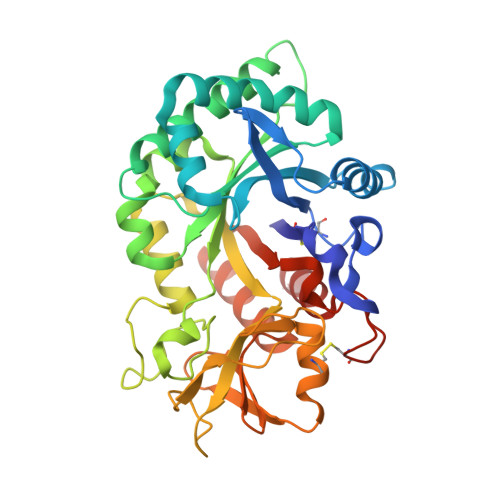
Crystal structure and carbohydrate-binding properties of the human cartilage glycoprotein-39
Fusetti, F., Pijning, T., Kalk, K.H., Bos, E., Dijkstra, B.W.(2003) J Biological Chem 278: 37753-37760
- PubMed: 12851408 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M303137200
- Primary Citation Related Structures:
1NWR, 1NWS, 1NWT, 1NWU - PubMed Abstract:
The human cartilage glycoprotein-39 (HCgp-39 or YKL40) is expressed by synovial cells and macrophages during inflammation. Its precise physiological role is unknown. However, it has been proposed that HCgp-39 acts as an autoantigen in rheumatoid arthritis, and high expression levels have been associated with cancer development. HCgp-39 shares high sequence homology with family 18 chitinases, and although it binds to chitin it lacks enzymatic activity. The crystal structure of HCgp-39 shows that the protein displays a (beta/alpha)8-barrel fold with an insertion of an alpha + beta domain. A 43-A long carbohydrate-binding cleft is present at the C-terminal side of the beta-strands in the (beta/alpha)8 barrel. Binding of chitin fragments of different lengths identified nine sugar-binding subsites in the groove. Protein-carbohydrate interactions are mainly mediated by stacking of side chains of aromatic amino acid residues. Surprisingly, the specificity of chitin binding to HCgp-39 depends on the length of the oligosaccharide. Although chitin disaccharides tend to occupy the distal subsites, longer chains bind preferably to the central subsites in the groove. Despite the absence of enzymatic activity, long chitin fragments are distorted upon binding, with the GlcNAc at subsite -1 in a boat conformation, similar to what has been observed in chitinases. The presence of chitin in the human body has never been documented so far. However, the binding features observed in the complex structures suggest that either chitin or a closely related oligosaccharide could act as the physiological ligand for HCgp-39.
- Laboratory of Biophysical Chemistry, Department of Chemistry, University of Groningen, Nijenborgh 4, 9747AG Groningen, The Netherlands.
Organizational Affiliation: