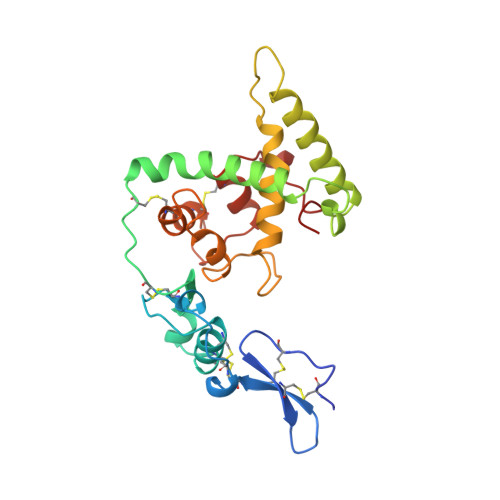
Crystal structure and mapping by site-directed mutagenesis of the collagen-binding epitope of an activated form of BM-40/SPARC/osteonectin.
Sasaki, T., Hohenester, E., Gohring, W., Timpl, R.(1998) EMBO J 17: 1625-1634
- PubMed: 9501084 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/17.6.1625
- Primary Citation Related Structures:
1NUB - PubMed Abstract:
The extracellular calcium-binding domain (positions 138-286) of the matrix protein BM-40 possesses a binding epitope of moderate affinity for several collagen types. This epitope was predicted to reside in helix alphaA and to be partially masked by helix alphaC. Here we show that deletion of helix alphaC produces a 10-fold increase in collagen affinity similar to that seen after proteolytic cleavage of this helix. The predicted removal of the steric constraint was clearly demonstrated by the crystal structure of the mutant at 2.8 A resolution. This constitutively activated mutant was used to map the collagen-binding site following alanine mutagenesis at 13 positions. Five residues were crucial for binding, R149 and N156 in helix alphaA, and L242, M245 and E246 in a loop region connecting the two EF hands of BM-40. These residues are spatially close and form a flat ring of 15 A diameter which matches the diameter of a triple-helical collagen domain. The mutations showed similar effects on binding to collagens I and IV, indicating nearly identical binding sites on both collagens. Selected mutations in the non-activated mutant DeltaI also reduced collagen binding, consistent with the same location of the epitope but in a more cryptic form in intact BM-40.
- Max-Planck-Institut für Biochemie, D-82152, Martinsried, Germany.
Organizational Affiliation: