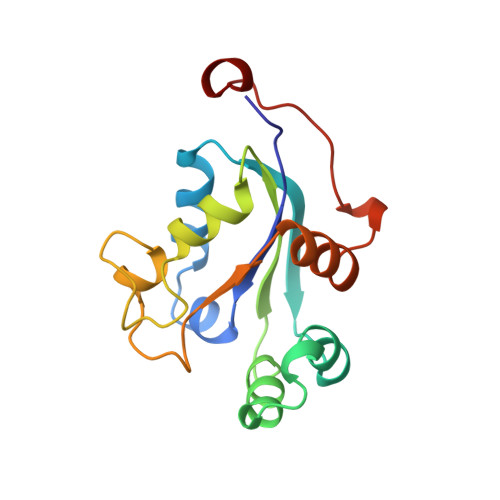
The crystal structure of a human nucleoside diphosphate kinase, NM23-H2.
Webb, P.A., Perisic, O., Mendola, C.E., Backer, J.M., Williams, R.L.(1995) J Mol Biology 251: 574-587
- PubMed: 7658474 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1995.0457
- Primary Citation Related Structures:
1NSK - PubMed Abstract:
The 2.8 A resolution X-ray structure of NM23-H2 has been determined by molecular replacement using the structure of Myxococcus xanthus nucleoside diphosphate (NDP) kinase. NM23-H2 is a human NDP kinase. The enzyme catalyses phosphoryl transfer, binds DNA, and can activate the transcription of the c-myc oncogene in vitro. NM23 has also been reported to be a suppressor of metastasis in some types of tumours. Whereas the M. xanthus NDP kinase is a tetramer, NM23-H2 is a hexamer. The fold of NM23-H2 is identical to the fold of other NDP kinases. Two antiparallel helices joined by a turn form one edge of the nucleotide binding cleft. This region moves in a hinge-like fashion in response to substrate binding and crystal packing forces. Additional differences in conformation among the NDP kinases are principally in regions involved in protein-protein contacts within the oligomers. The only protein-protein interaction conserved among all NDP kinases is a dimeric interaction. Several mutations of NM23-H2 have been detected in tumour tissues. These mutations do not involve residues interacting with the substrates, and probably destabilise the enzyme without directly affecting the catalytic activity. Low level phosphorylation of serines has been reported for NM23 both in vitro and in vivo. The structure of the hexamer indicates that two serine residues that have been reported as being phosphorylated, Ser44 and Ser122, are on the surface of the hexamer, and are likely to be phosphorylated by exogenous kinases. In contrast, Ser120 is buried, and is most likely phosphorylated by a direct transfer from the phosphohistidine intermediate of the reaction mechanism.
- Centre for Protein Engineering, MRC Centre, Cambridge, UK.
Organizational Affiliation: