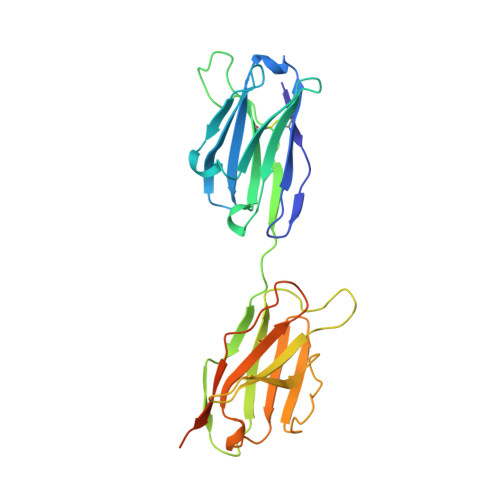
The 2.0-A resolution crystal structure of a trimeric antibody fragment with noncognate VH-VL domain pairs shows a rearrangement of VH CDR3.
Pei, X.Y., Holliger, P., Murzin, A.G., Williams, R.L.(1997) Proc Natl Acad Sci U S A 94: 9637-9642
- PubMed: 9275175 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.94.18.9637
- Primary Citation Related Structures:
1NQB - PubMed Abstract:
The 2.0-A resolution x-ray crystal structure of a novel trimeric antibody fragment, a "triabody," has been determined. The trimer is made up of polypeptides constructed in a manner identical to that previously described for some "diabodies": a VL domain directly fused to the C terminus of a VH domain-i.e., without any linker sequence. The trimer has three Fv heads with the polypeptides arranged in a cyclic, head-to-tail fashion. For the particular structure reported here, the polypeptide was constructed with a VH domain from one antibody fused to the VL domain from an unrelated antibody giving rise to "combinatorial" Fvs upon formation of the trimer. The structure shows that the exchange of the VL domain from antibody B1-8, a Vlambda domain, with the VL domain from antibody NQ11, a Vkappa domain, leads to a dramatic conformational change in the VH CDR3 loop of antibody B1-8. The magnitude of this change is similar to the largest of the conformational changes observed in antibody fragments in response to antigen binding. Combinatorial pairing of VH and VL domains constitutes a major component of antibody diversity. Conformationally flexible antigen-binding sites capable of adapting to the specific CDR3 loop context created upon VH-VL pairing may be employed by the immune system to maximize the structural diversity of the immune response.
- Centre for Protein Engineering, Medical Research Council Centre, Hills Road, Cambridge CB2 2QH, United Kingdom.
Organizational Affiliation: