Structural basis for endothelial nitric oxide synthase binding to calmodulin
Aoyagi, M., Arvai, A.S., Tainer, J.A., Getzoff, E.D.(2003) EMBO J 22: 766-775
- PubMed: 12574113 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/cdg078
- Primary Citation Related Structures:
1NIW - PubMed Abstract:
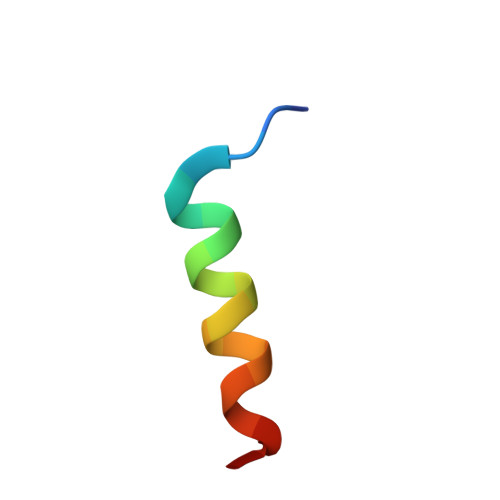
The enzyme nitric oxide synthase (NOS) is exquisitely regulated in vivo by the Ca(2+) sensor protein calmodulin (CaM) to control production of NO, a key signaling molecule and cytotoxin. The differential activation of NOS isozymes by CaM has remained enigmatic, despite extensive research. Here, the crystallographic structure of Ca(2+)-loaded CaM bound to a 20 residue peptide comprising the endothelial NOS (eNOS) CaM-binding region establishes their individual conformations and intermolecular interactions, and suggests the basis for isozyme-specific differences. The alpha-helical eNOS peptide binds in an antiparallel orientation to CaM through extensive hydrophobic interactions. Unique NOS interactions occur with: (i). the CaM flexible central linker, explaining its importance in NOS activation; and (ii). the CaM C-terminus, explaining the NOS-specific requirement for a bulky, hydrophobic residue at position 144. This binding mode expands mechanisms for CaM-mediated activation, explains eNOS deactivation by Thr495 phosphorylation, and implicates specific hydrophobic residues in the Ca(2+) independence of inducible NOS.
- Department of Molecular Biology and The Skaggs Institute for Chemical Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: