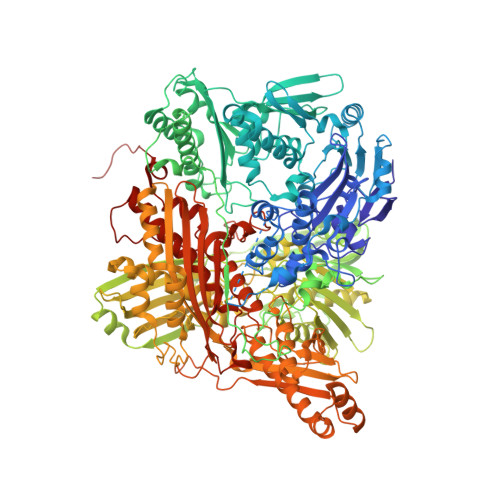
An Extremely Potent Inhibitor of Xanthine Oxidoreductase: Crystal Structure of the Enzyme-Inhibitor Complex and Mechanism of Inhibition
Okamoto, K., Eger, B.T., Nishino, T., Kondo, S., Pai, E.F., Nishino, T.(2003) J Biological Chem 278: 1848-1855
- PubMed: 12421831 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M208307200
- Primary Citation Related Structures:
1N5X - PubMed Abstract:
TEI-6720 (2-(3-cyano-4-isobutoxyphenyl)-4-methyl-5-thiazolecarboxylic acid) is an extremely potent inhibitor of xanthine oxidoreductase. Steady state kinetics measurements exhibit mixed type inhibition with K(i) and K(i)' values of 1.2 +/- 0.05 x 10(-10) m and 9 +/- 0.05 x 10(-10) m, respectively. Fluorescence-monitored titration experiments showed that TEI-6720 bound very tightly to both the active and the inactive desulfo-form of the enzyme. The dissociation constant determined for the desulfo-form was 2 +/- 0.03 x 10(-9) m; for the active form, the corresponding number was too low to allow accurate measurements. The crystal structure of the active sulfo-form of milk xanthine dehydrogenase complexed with TEI-6720 and determined at 2.8-A resolution revealed the inhibitor molecule bound in a long, narrow channel leading to the molybdenum-pterin active site of the enzyme. It filled up most of the channel and the immediate environment of the cofactor, very effectively inhibiting the activity of the enzyme through the prevention of substrate binding. Although the inhibitor did not directly coordinate to the molybdenum ion, numerous hydrogen bonds as well as hydrophobic interactions with the protein matrix were observed, most of which are also used in substrate recognition.
- Department of Biochemistry and Molecular Biology, Nippon Medical School, 1-1-5 Sendagi, Bunkyo-ku, Tokyo 113-0022, Japan.
Organizational Affiliation: