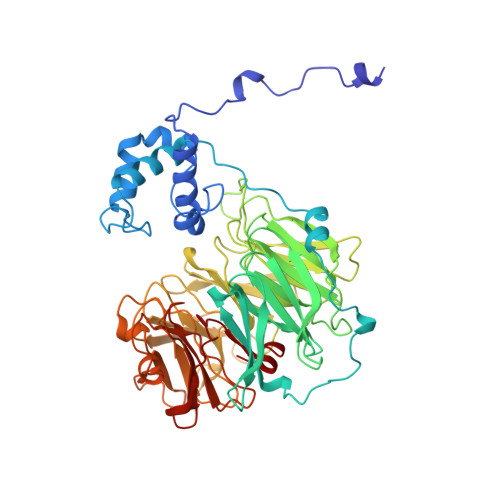
Does the reduction of c heme trigger the conformational change of crystalline nitrite reductase?
Nurizzo, D., Cutruzzola, F., Arese, M., Bourgeois, D., Brunori, M., Cambillau, C., Tegoni, M.(1999) J Biological Chem 274: 14997-15004
- PubMed: 10329702 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.274.21.14997
- Primary Citation Related Structures:
1N15, 1N50, 1N90 - PubMed Abstract:
The structures of nitrite reductase from Paracoccus denitrificans GB17 (NiR-Pd) and Pseudomonas aeruginosa (NiR-Pa) have been described for the oxidized and reduced state (Fülöp, V., Moir, J. W. B., Ferguson, S. J., and Hajdu, J. (1995) Cell 81, 369-377; Nurizzo, D., Silvestrini, M. C., Mathieu, M., Cutruzzolà, F., Bourgeois, D., Fülöp, V., Hajdu, J., Brunori, M., Tegoni, M., and Cambillau, C. (1997) Structure 5, 1157-1171; Nurizzo, D., Cutruzzolà, F., Arese, M., Bourgeois, D., Brunori, M., Cambillau, C. , and Tegoni, M. (1998) Biochemistry 37, 13987-13996). Major conformational rearrangements are observed in the extreme states although they are more substantial in NiR-Pd. The four structures differ significantly in the c heme domains. Upon reduction, a His17/Met106 heme-ligand switch is observed in NiR-Pd together with concerted movements of the Tyr in the distal site of the d1 heme (Tyr10 in NiR-Pa, Tyr25 in NiR-Pd) and of a loop of the c heme domain (56-62 in NiR-Pa, 99-116 in NiR-Pd). Whether the reduction of the c heme, which undergoes the major rearrangements, is the trigger of these movements is the question addressed by our study. This conformational reorganization is not observed in the partially reduced species, in which the c heme is partially or largely (15-90%) reduced but the d1 heme is still oxidized. These results suggest that the d1 heme reduction is likely to be responsible of the movements. We speculate about the mechanistic explanation as to why the opening of the d1 heme distal pocket only occurs upon electron transfer to the d1 heme itself, to allow binding of the physiological substrate NO2- exclusively to the reduced metal center.
- Architecture et Fonction des Macromolécules Biologiques, UPR9039-CNRS, IBSM, 31, Ch. Joseph Aiguier, Marseille Cedex 20, France.
Organizational Affiliation: