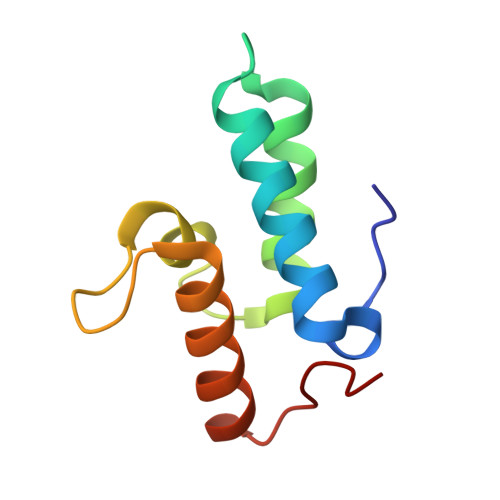
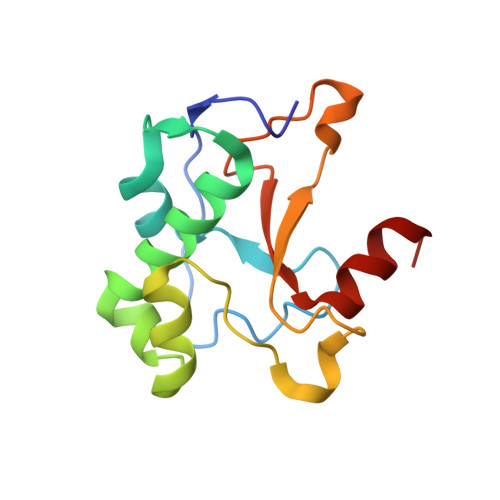
Metal ions and phosphate binding in the H-N-H motif: crystal structures of the nuclease domain of ColE7/Im7 in complex with a phosphate ion and different divalent metal ions
Sui, M.J., Tsai, L.C., Hsia, K.C., Doudeva, L.G., Ku, W.Y., Han, G.W., Yuan, H.S.(2002) Protein Sci 11: 2947-2957
- PubMed: 12441392 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.0220602
- Primary Citation Related Structures:
1MZ8 - PubMed Abstract:
H-N-H is a motif found in the nuclease domain of a subfamily of bacteria toxins, including colicin E7, that are capable of cleaving DNA nonspecifically. This H-N-H motif has also been identified in a subfamily of homing endonucleases, which cleave DNA site specifically. To better understand the role of metal ions in the H-N-H motif during DNA hydrolysis, we crystallized the nuclease domain of colicin E7 (nuclease-ColE7) in complex with its inhibitor Im7 in two different crystal forms, and we resolved the structures of EDTA-treated, Zn(2+)-bound and Mn(2+)-bound complexes in the presence of phosphate ions at resolutions of 2.6 A to 2.0 A. This study offers the first determination of the structure of a metal-free and substrate-free enzyme in the H-N-H family. The H-N-H motif contains two antiparallel beta-strands linked to a C-terminal alpha-helix, with a divalent metal ion located in the center. Here we show that the metal-binding sites in the center of the H-N-H motif, for the EDTA-treated and Mg(2+)-soaked complex crystals, were occupied by water molecules, indicating that an alkaline earth metal ion does not reside in the same position as a transition metal ion in the H-N-H motif. However, a Zn(2+) or Mn(2+) ions were observed in the center of the H-N-H motif in cases of Zn(2+) or Mn(2+)-soaked crystals, as confirmed in anomalous difference maps. A phosphate ion was found to bridge between the divalent transition metal ion and His545. Based on these structures and structural comparisons with other nucleases, we suggest a functional role for the divalent transition metal ion in the H-N-H motif in stabilizing the phosphoanion in the transition state during hydrolysis.
- Graduate Institute of Life Science, National Defense Medical Center, Taipei, Taiwan 11472, ROC.
Organizational Affiliation: