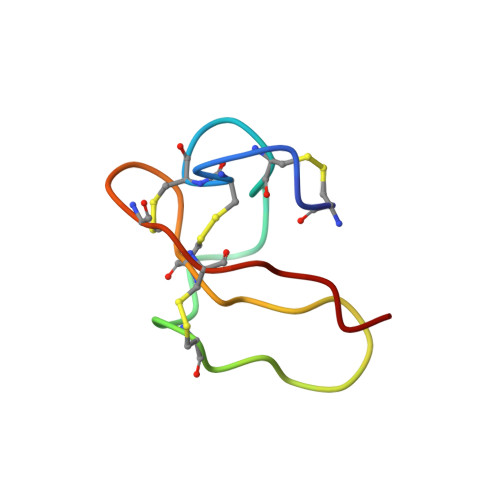
NMR Solution Structure of the Non-RGD Disintegrin Obtustatin
Moreno-Murciano, M.P., Monleon, D., Marcinkiewicz, C., Calvete, J.J., Celda, B.(2003) J Mol Biology 329: 135-145
- PubMed: 12742023 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(03)00371-1
- Primary Citation Related Structures:
1MPZ - PubMed Abstract:
The solution structure of obtustatin, a novel non-RGD disintegrin of 41 residues isolated from Vipera lebetina obtusa venom, and a potent and selective inhibitor of the adhesion of integrin alpha(1)beta(1) to collagen IV, has been determined by two-dimensional nuclear magnetic resonance. Almost the whole set of chemical shifts for 1H, 13C and 15N were assigned at natural abundance from 2D homonuclear and heteronuclear 500 MHz, 600 MHz and 800 MHz spectra at pH 3.0 recorded at 298 K and 303 K. Final structural constraints consisted of 302 non-redundant NOE (95 long-range, 60 medium, 91 sequential and 56 intra-residue), four disulfide bond distances, five chi1 dihedral angles and four hydrogen bonds. The 20 conformers with lowest total energy had no NOE violations greater than 0.35A or dihedral angle violations greater than 12 degrees. The average root-mean-square deviation (RMSD) for backbone atoms of all residues among the 20 conformers was 1.1A and 0.6A for the 29 best-defined residues. Obtustatin lacks any secondary structure. Compared to all known disintegrin structures in which the RGD motif is located at the apex of an 11 residue hairpin loop, the active KTS tripeptide of obtustatin is oriented towards a side of its nine residue integrin-binding loop. The C-terminal tail is near to the active loop, and these two structural elements display the largest atomic displacements due to local conformational disorder. Double cross-peaks for W20, Y28 and H27 in the aromatic region of TOCSY spectra, local RMSD values for these residues, and positive cross-peaks in a ROESY spectrum (600 MHz, 100 ms mixing time), suggest that these residues act as a hinge allowing for the overall flexibility of the entire integrin-binding loop. These distinct structural features, along with its different electrostatic surface potential in relation to other known disintegrins, may confer to obtustatin its reported alpha(1)beta(1) integrin inhibitory selectivity.
- Departamento de Qui;mica Fi;sica, Universitat de València, c/Dr Moliner, 50, 46100 Burjassot, Valencia, Spain.
Organizational Affiliation: