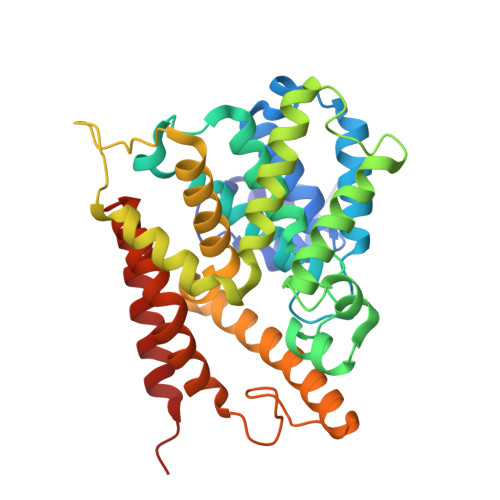
Crystal structure of phosphodiesterase 4D and inhibitor complex
Lee, M.E., Markowitz, J., Lee, J.-O., Lee, H.(2002) FEBS Lett 530: 53-58
- PubMed: 12387865 Search on PubMed
- DOI: https://doi.org/10.1016/s0014-5793(02)03396-3
- Primary Citation Related Structures:
1MKD - PubMed Abstract:
Cyclic nucleotide phosphodiesterases (PDEs) regulate physiological processes by degrading intracellular second messengers, adenosine-3',5'-cyclic phosphate or guanosine-3',5'-cyclic phosphate. The first crystal structure of PDE4D catalytic domain and a bound inhibitor, zardaverine, was determined. Zardaverine binds to a highly conserved pocket that includes the catalytic metal binding site. Zardaverine fills only a portion of the active site pocket. More selective PDE4 inhibitors including rolipram, cilomilast and roflumilast have additional functional groups that can utilize the remaining empty space for increased binding energy and selectivity. In the crystal structure, the catalytic domain of PDE4D possesses an extensive dimerization interface containing residues that are highly conserved in PDE1, 3, 4, 8 and 9. Mutations of R358D or D322R among these interface residues prohibit dimerization of the PDE4D catalytic domain in solution.
- Department of Chemistry, Korea Advanced Institute of Science and Technology, 373-1 Kusong-dong, Yusong-gu, Daejon 305-701, South Korea.
Organizational Affiliation: