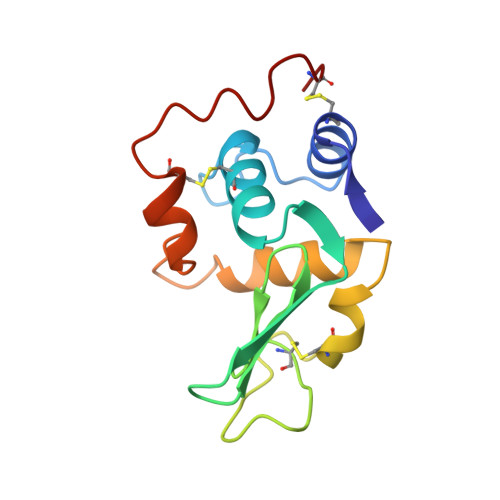
Enthalpic destabilization of a mutant human lysozyme lacking a disulfide bridge between cysteine-77 and cysteine-95.
Kuroki, R., Inaka, K., Taniyama, Y., Kidokoro, S., Matsushima, M., Kikuchi, M., Yutani, K.(1992) Biochemistry 31: 8323-8328
- PubMed: 1525170
- DOI: https://doi.org/10.1021/bi00150a028
- Primary Citation Related Structures:
1LZ4 - PubMed Abstract:
To understand the role of disulfide bridges in protein stability, the thermodynamic changes in the denaturation of two mutant human lysozymes lacking a disulfide bridge between Cys-77 and Cys-95 (C77A and C77/95A) were analyzed using differential scanning calorimetry (DSC). At pH 3.0 and 57 degrees C, the stabilities of both the C77A and C77/95A mutants were decreased about 4.6 kcal.mol-1 in Gibbs free energy change. Under the same conditions, the enthalpy changes (delta H) were 94.8 and 90.8 kcal.mol-1, respectively, which were smaller than that of the wild type (100.8 kcal.mol-1). The destabilization of the mutants was caused by enthalpic factors. Although X-ray crystallography indicated that the mutants preserve the wild-type tertiary structure, removal of the disulfide bridge increased the flexibility of the native state of the mutants. This was indicated both by an increase in the crystallographic thermal factors (B-factors) and by a decrease in the affinity of N-acetylglucosamine trimer [(NAG)3] observed using isothermal titration calorimetry (DTC) due to entropic effects. Thus, the effect of cross-linking on the stability of a protein is not solely explained by the entropy change in denaturation.
- Protein Engineering Research Institute, Osaka, Japan.
Organizational Affiliation: