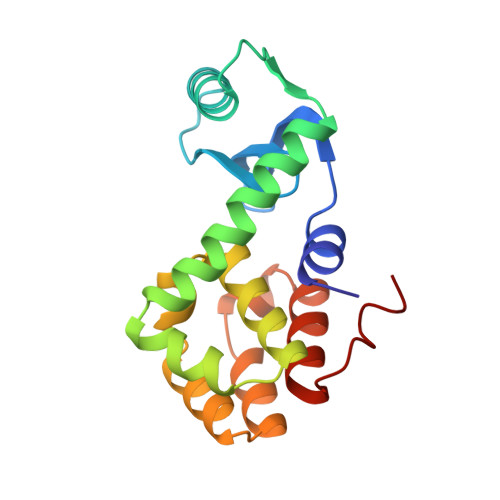
Crystal structure of T4-lysozyme generated from synthetic coding DNA expressed in Escherichia coli.
Rose, D.R., Phipps, J., Michniewicz, J., Birnbaum, G.I., Ahmed, F.R., Muir, A., Anderson, W.F., Narang, S.(1988) Protein Eng 2: 277-282
- PubMed: 3074306 Search on PubMed
- DOI: https://doi.org/10.1093/protein/2.4.277
- Primary Citation Related Structures:
1LYD - PubMed Abstract:
The polypeptide produced by expressing a chemically synthesized gene coding for the amino-acid sequence of T4-lysozyme has been crystallized and subjected to X-ray diffraction. The crystal structure has been refined to a standard R-factor of 0.191 for data between 8 and 2 A resolution. The refined model is essentially the same as the well-known structure of wild-type T4-lysozyme determined previously by Matthews et al. (1987). Some small changes in the C-terminal region, which is important in maintaining the folded structure, have been noted. In addition to confirming that the synthetic gene product is very close to the wild type, this structure provides a benchmark for protein engineering experiments on the folding and the catalytic activity of this molecule by the method of gene synthesis.
- Division of Biological Sciences, National Research Council of Canada, Ottawa.
Organizational Affiliation: