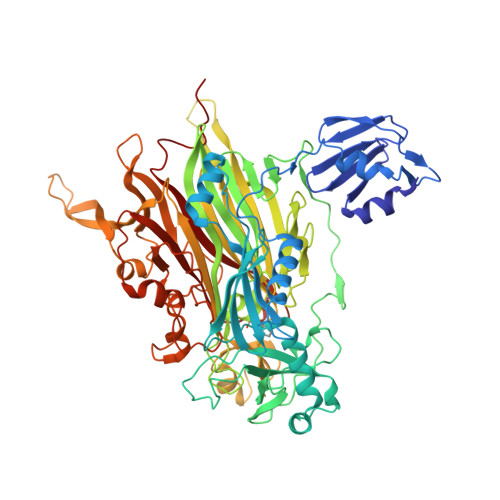
Medical implications from the crystal structure of a copper-containing amine oxidase complexed with the antidepressant drug tranylcypromine.
Wilmot, C.M., Saysell, C.G., Blessington, A., Conn, D.A., Kurtis, C.R., McPherson, M.J., Knowles, P.F., Phillips, S.E.(2004) FEBS Lett 576: 301-305
- PubMed: 15498552 Search on PubMed
- DOI: https://doi.org/10.1016/j.febslet.2004.09.031
- Primary Citation Related Structures:
1LVN - PubMed Abstract:
The X-ray crystal structure of the copper-containing quinoprotein amine oxidase from E. coli has been determined in complex with the antidepressant drug tranylcypromine to 2.4 A resolution. The drug is a racemic mix of two enantiomers, but only one is seen bound to the enzyme. The other enantiomer is not acting as a substrate for the enzyme as no catalytic activity was detected when the enzyme was initially exposed to the drug. The inhibition of human copper amine oxidases could be a source of side-effects in its use as an antidepressant to inhibit the flavin-containing monoamine oxidases in the brain.
- Astbury Centre for Structural Molecular Biology, School of Biochemistry and Molecular Biology, University of Leeds, Leeds LS2 9JT, UK. wilmo004@umn.edu
Organizational Affiliation: