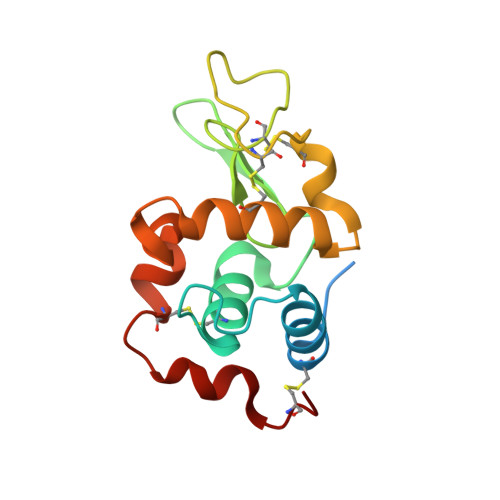
Crystal structure of the mutant D52S hen egg white lysozyme with an oligosaccharide product.
Hadfield, A.T., Harvey, D.J., Archer, D.B., MacKenzie, D.A., Jeenes, D.J., Radford, S.E., Lowe, G., Dobson, C.M., Johnson, L.N.(1994) J Mol Biology 243: 856-872
- PubMed: 7966306 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1994.1688
- Primary Citation Related Structures:
1LSY, 1LSZ - PubMed Abstract:
The crystal structure of a mutant hen egg white lysozyme, in which the key catalytic residue aspartic acid 52 has been changed to a serine residue (D52S HEWL), has been determined and refined to a crystallographic R value of 0.173 for all data F > 0 between 8 and 1.9 A resolution. The D52S HEWL structure is very similar to the native HEWL structure (r.m.s. deviation of main-chain atoms 0.20 A). Small shifts that result from the change in hydrogen bonding pattern on substitution of Asp by Ser were observed in the loop between beta-strands in the region of residues 46 to 49. D52S HEWL exhibits less than 1% activity against the bacterial cell wall substrate. Cocrystallisation experiments with the hexasaccharide substrate beta(1-4) polymer of N-acetyl-D-glucosamine (GlcNAc6) resulted in crystals between 5 days and 14 days after the initial mixing of enzyme and substrate. Analysis by laser absorption mass spectrometry of the oligosaccharides present after incubation with native and D52S HEWL under conditions similar to those used for crystal growth showed that after 14 days with native HEWL complete catalysis to GlcNAc3. GlcNAc2 and GlcNac had occurred but with D52S HEWL only partial catalysis to the major products GlcNAc4 and GlcNAc2 had occurred and at least 50% of the GlcNAc6 remained intact. X-ray analysis of the D52S-oligosaccharide complex crystals showed that they contained the product GlcNAc4. The structure of the D52S HEWL-GlcNAc4 complex has been determined and refined to an R value of 0.160 for data between 8 and 2 A resolution. GlcNAc4 occupies sites A to D in the active site cleft. Careful refinement and examination of 2Fo-Fc electron density maps showed that the sugar in site D has the sofa conformation, a conformation previously observed with the HEWL complex with tetra-N-acetylglucosamine lactone transition state analogue, the HEWL complex with the cell wall trisaccharide and the phage T4 lysozyme complex with a cell wall product. The semi-axial C(5)-C(6) geometry of the sofa is stabilised by hydrogen bonds from the O-6 hydroxyl group to the main-chain N of Val109 and main-chain O of Ala107. The sugar in site D adopts the alpha configuration, seemingly in conflict with the observation that the hydrolysis of beta (1-4) glycosidie linkage by HEWL proceeds with 99.9% retention of beta-configuration.(ABSTRACT TRUNCATED AT 400 WORDS)
- Oxford Centre for Molecular Sciences, University of Oxford, U.K.
Organizational Affiliation: