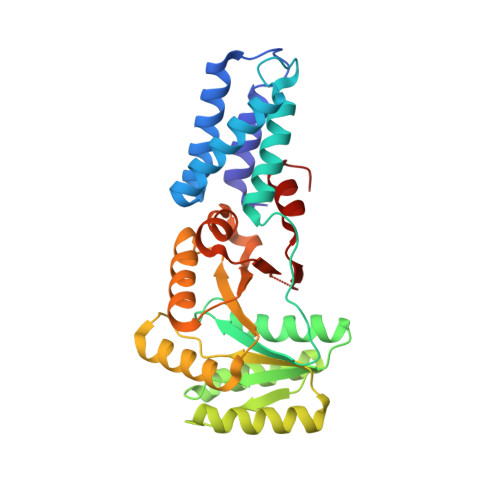
Structural Basis for Mobility in the 1.1 A Crystal Structure of the NG Domain of Thermus aquaticus Ffh
Ramirez, U.D., Minasov, G., Focia, P.J., Stroud, R.M., Walter, P., Kuhn, P., Freymann, D.M.(2002) J Mol Biology 320: 783-799
- PubMed: 12095255 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/s0022-2836(02)00476-x
- Primary Citation Related Structures:
1LS1 - PubMed Abstract:
The NG domain of the prokaryotic signal recognition protein Ffh is a two-domain GTPase that comprises part of the prokaryotic signal recognition particle (SRP) that functions in co-translational targeting of proteins to the membrane. The interface between the N and G domains includes two highly conserved sequence motifs and is adjacent in sequence and structure to one of the conserved GTPase signature motifs. Previous structural studies have shown that the relative orientation of the two domains is dynamic. The N domain of Ffh has been proposed to function in regulating the nucleotide-binding interactions of the G domain. However, biochemical studies suggest a more complex role for the domain in integrating communication between signal sequence recognition and interaction with receptor. Here, we report the structure of the apo NG GTPase of Ffh from Thermus aquaticus refined at 1.10 A resolution. Although the G domain is very well ordered in this structure, the N domain is less well ordered, reflecting the dynamic relationship between the two domains previously inferred. We demonstrate that the anisotropic displacement parameters directly visualize the underlying mobility between the two domains, and present a detailed structural analysis of the packing of the residues, including the critical alpha4 helix, that comprise the interface. Our data allows us to propose a structural explanation for the functional significance of sequence elements conserved at the N/G interface.
- Department of Molecular Pharmacology and Biological Chemistry, Northwestern University Medical School, 303 E Chicago Avenue, MC S215, Chicago, IL 60611-3008, USA.
Organizational Affiliation: