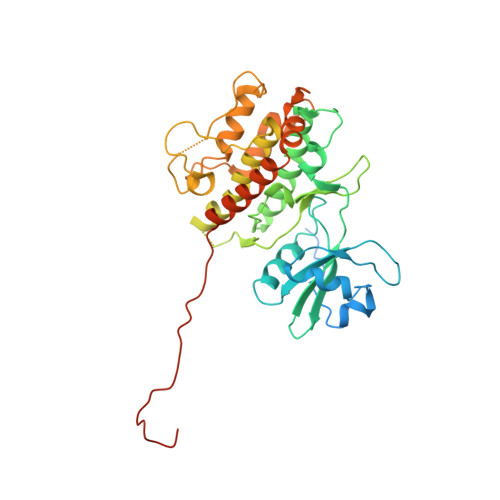
Structure of Mitogen-activated Protein Kinase-activated Protein (MAPKAP) Kinase 2 Suggests a Bifunctional Switch That Couples Kinase Activation with Nuclear Export
Meng, W., Swenson, L.L., Fitzgibbon, M.J., Hayakawa, K., Ter Haar, E., Behrens, A.E., Fulghum, J.R., Lippke, J.A.(2002) J Biological Chem 277: 37401-37405
- PubMed: 12171911 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.C200418200
- Primary Citation Related Structures:
1KWP - PubMed Abstract:
MAPK-activated protein kinase 2 (MAPKAPK2), one of several kinases directly phosphorylated and activated by p38 MAPK, plays a central role in the inflammatory response. The activated MAPKAPK2 phosphorylates its nuclear targets CREB/ATF1, serum response factor, and E2A protein E47 and its cytoplasmic targets HSP25/27, LSP-1, 5-lipoxygenase, glycogen synthase, and tyrosine hydroxylase. The crystal structure of unphosphorylated MAPKAPK2, determined at 2.8 A resolution, includes the kinase domain and the C-terminal regulatory domain. Although the protein is inactive, the kinase domain adopts an active conformation with aspartate 366 mimicking the missing phosphorylated threonine 222 in the activation loop. The C-terminal regulatory domain forms a helix-turn-helix plus a long strand. Phosphorylation of threonine 334, which is located between the kinase domain and the C-terminal regulatory domain, may serve as a switch for MAPKAPK2 nuclear import and export. Phosphorylated MAPKAPK2 masks the nuclear localization signal at its C terminus by binding to p38. It unmasks the nuclear export signal, which is part of the second C-terminal helix packed along the surface of kinase domain C-lobe, and thereby carries p38 to the cytoplasm.
- Vertex Pharmaceuticals Inc., Cambridge, Massachusetts 02139, USA. wuyi_meng@vpharm.com
Organizational Affiliation: