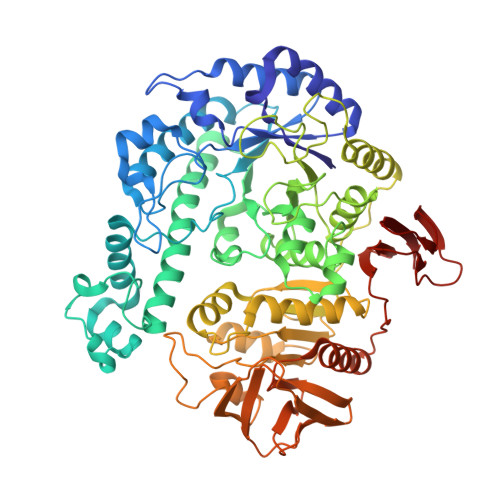
Trimeric Crystal Structure of the Glycoside Hydrolase Family 42 beta-Galactosidase from Thermus thermophilus A4 and the Structure of its Complex with Galactose
Hidaka, M., Fushinobu, S., Ohtsu, N., Motoshima, H., Matsuzawa, H., Shoun, H., Wakagi, T.(2002) J Mol Biology 322: 79-91
- PubMed: 12215416 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(02)00746-5
- Primary Citation Related Structures:
1KWG, 1KWK - PubMed Abstract:
The beta-galactosidase from an extreme thermophile, Thermus thermophilus A4 (A4-beta-Gal), is thermostable and belongs to the glycoside hydrolase family 42 (GH-42). As the first known structures of a GH-42 enzyme, we determined the crystal structures of free and galactose-bound A4-beta-Gal at 1.6A and 2.2A resolution, respectively. A4-beta-Gal forms a homotrimeric structure resembling a flowerpot. Each monomer has an active site located inside a large central tunnel. The N-terminal domain of A4-beta-Gal has a TIM barrel fold, as predicted from hydrophobic cluster analysis. The putative catalytic residues of A4-beta-Gal (Glu141 and Glu312) superimpose well with the catalytic residues of Escherichia coli beta-galactosidase. The environment around the catalytic nucleophile (Glu312) is similar to that in the case of E.coli beta-galactosidase, but the recognition mechanism for a substrate is different. Trp182 of the next subunit of the trimer constitutes a part of the active-site pocket, indicating that the trimeric structure is essential for the enzyme activity. Structural comparison with other glycoside hydrolases revealed that many features of the 4/7 superfamily are conserved in the A4-beta-Gal structure. On the basis of the results of 1H NMR spectroscopy, A4-beta-Gal was determined to be a "retaining" enzyme. Interestingly, the active site was similar with those of retaining enzymes, but the overall fold of the TIM barrel domain was very similar to that of an inverting enzyme, beta-amylase.
- Department of Biotechnology, The University of Tokyo, Japan.
Organizational Affiliation: