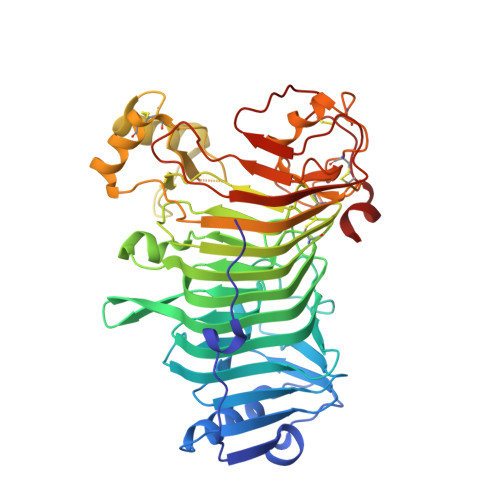
The Structural Bases of the Processive Degradation of iota-Carrageenan, a Main Cell Wall Polysaccharide of Red Algae.
Michel, G., Helbert, W., Kahn, R., Dideberg, O., Kloareg, B.(2003) J Mol Biology 334: 421-433
- PubMed: 14623184 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2003.09.056
- Primary Citation Related Structures:
1KTW - PubMed Abstract:
iota-Carrageenans are sulfated 1,3-alpha-1,4-beta-galactans from the cell walls of red algae, which auto-associate into crystalline fibers made of aggregates of double-stranded helices. iota-Carrageenases, which constitute family 82 of glycoside hydrolases, fold into a right-handed beta-helix. Here, the structure of Alteromonas fortis iota-carrageenase bound to iota-carrageenan fragments was solved at 2.0A resolution (PDB 1KTW). The enzyme holds a iota-carrageenan tetrasaccharide (subsites +1 to +4) and a disaccharide (subsites -3, -4), thus providing the first direct determination of a 3D structure of iota-carrageenan. Electrostatic interactions between basic protein residues and the sulfate substituents of the polysaccharide chain dominate iota-carrageenan recognition. Glu245 and Asp247 are the proton donor and the base catalyst, respectively. C-terminal domain A, which was highly flexible in the native enzyme structure, adopts a alpha/beta-fold, also found in DNA/RNA-binding domains. In the substrate-enzyme complex, this polyanion-binding module shifts toward the beta-helix groove, forming a tunnel. Thus, from an open conformation which allows for the initial endo-attack of iota-carrageenan chains, the enzyme switches to a closed-tunnel form, consistent with its highly processive character, as seen from the electron-microscopy analysis of the degradation of iota-carrageenan fibers.
- Végétaux Marins et Biomolécules, UMR 7139 (CNRS/UPMC/Laboratories Goëmar), Station Biologique de Roscoff, Place Georges Teissier, BP 74, 29682 Roscoff Cedex, Brittany, France.kloareg@sb-rosecoff.ff
Organizational Affiliation: