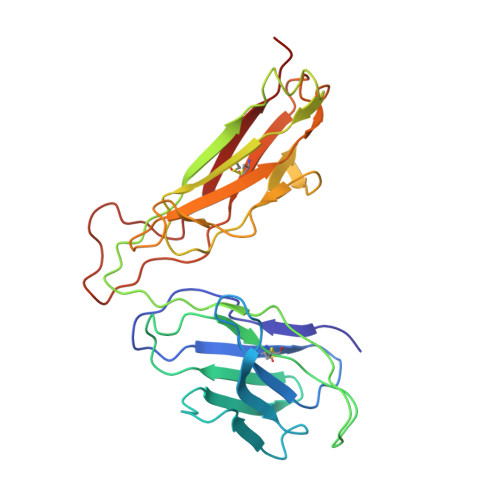
Structures of two streptococcal superantigens bound to TCR beta chains reveal diversity in the architecture of T cell signaling complexes.
Sundberg, E.J., Li, H., Llera, A.S., McCormick, J.K., Tormo, J., Schlievert, P.M., Karjalainen, K., Mariuzza, R.A.(2002) Structure 10: 687-699
- PubMed: 12015151 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(02)00759-1
- Primary Citation Related Structures:
1KTK, 1L0X, 1L0Y - PubMed Abstract:
Superantigens (SAGs) crosslink MHC class II and TCR molecules, resulting in an overstimulation of T cells associated with human disease. SAGs interact with several different surfaces on MHC molecules, necessitating the formation of multiple distinct MHC-SAG-TCR ternary signaling complexes. Variability in SAG-TCR binding modes could also contribute to the structural heterogeneity of SAG-dependent signaling complexes. We report crystal structures of the streptococcal SAGs SpeA and SpeC in complex with their corresponding TCR beta chain ligands that reveal distinct TCR binding modes. The SpeC-TCR beta chain complex structure, coupled with the recently determined SpeC-HLA-DR2a complex structure, provides a model for a novel T cell signaling complex that precludes direct TCR-MHC interactions. Thus, highly efficient T cell activation may be achieved through structurally diverse strategies of TCR ligation.
- Center for Advanced Research in Biotechnology, W.M. Keck Laboratory for Structural Biology, University of Maryland Biotechnology Institute, 9600 Gudelsky Drive, Rockville, MD 20850, USA.
Organizational Affiliation: