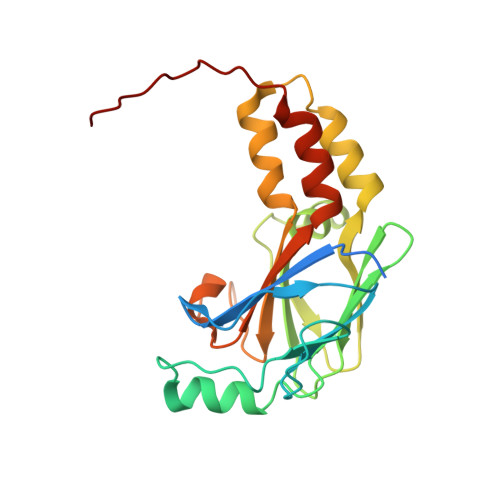
Structural basis of Smad1 activation by receptor kinase phosphorylation.
Qin, B.Y., Chacko, B.M., Lam, S.S., de Caestecker, M.P., Correia, J.J., Lin, K.(2001) Mol Cell 8: 1303-1312
- PubMed: 11779505 Search on PubMed
- DOI: https://doi.org/10.1016/s1097-2765(01)00417-8
- Primary Citation Related Structures:
1KHU - PubMed Abstract:
Phosphorylation of Smad1 at the conserved carboxyl terminal SVS sequence activates BMP signaling. Here we report the crystal structure of the Smad1 MH2 domain in a conformation that reveals the structural effects of phosphorylation and a molecular mechanism for activation. Within a trimeric subunit assembly, the SVS sequence docks near two putative phosphoserine binding pockets of the neighboring molecule, in a position ready to interact upon phosphorylation. The MH2 domain undergoes concerted conformational changes upon activation, which signal Smad1 dissociation from the receptor kinase for subsequent heteromeric assembly with Smad4. Biochemical and modeling studies reveal unique favorable interactions within the Smad1/Smad4 heteromeric interface, providing a structural basis for their association in signaling.
- Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, Worcester, MA 01655, USA.
Organizational Affiliation: