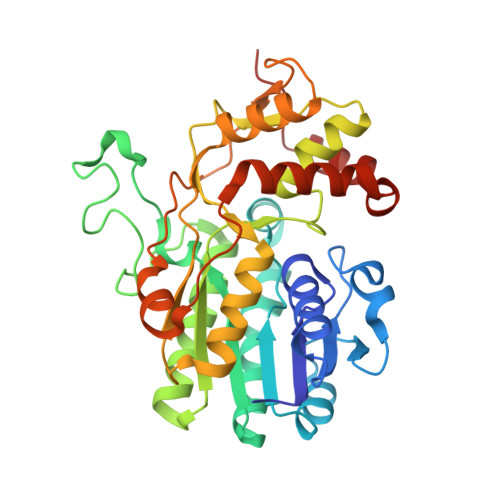
Toward a structural understanding of the dehydratase mechanism.
Allard, S.T., Beis, K., Giraud, M.F., Hegeman, A.D., Gross, J.W., Wilmouth, R.C., Whitfield, C., Graninger, M., Messner, P., Allen, A.G., Maskell, D.J., Naismith, J.H.(2002) Structure 10: 81-92
- PubMed: 11796113 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00694-3
- Primary Citation Related Structures:
1KEP, 1KER, 1KET, 1KEU, 1KEW - PubMed Abstract:
dTDP-D-glucose 4,6-dehydratase (RmlB) was first identified in the L-rhamnose biosynthetic pathway, where it catalyzes the conversion of dTDP-D-glucose into dTDP-4-keto-6-deoxy-D-glucose. The structures of RmlB from Salmonella enterica serovar Typhimurium in complex with substrate deoxythymidine 5'-diphospho-D-glucose (dTDP-D-glucose) and deoxythymidine 5'-diphosphate (dTDP), and RmlB from Streptococcus suis serotype 2 in complex with dTDP-D-glucose, dTDP, and deoxythymidine 5'-diphospho-D-pyrano-xylose (dTDP-xylose) have all been solved at resolutions between 1.8 A and 2.4 A. The structures show that the active sites are highly conserved. Importantly, the structures show that the active site tyrosine functions directly as the active site base, and an aspartic and glutamic acid pairing accomplishes the dehydration step of the enzyme mechanism. We conclude that the substrate is required to move within the active site to complete the catalytic cycle and that this movement is driven by the elimination of water. The results provide insight into members of the SDR superfamily.
- Centre for Biomolecular Sciences, North Haugh, The University, St. Andrews, Fife KY16 9ST, Scotland, United Kingdom.
Organizational Affiliation: