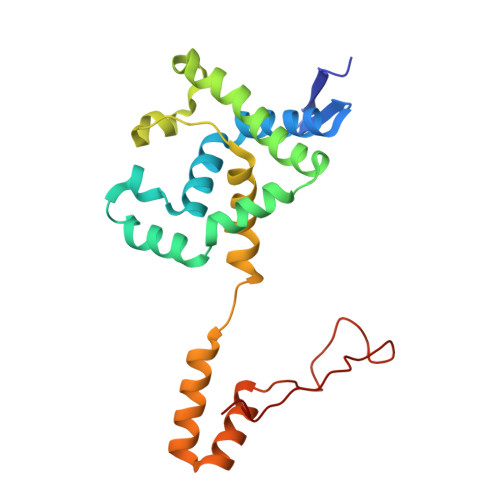
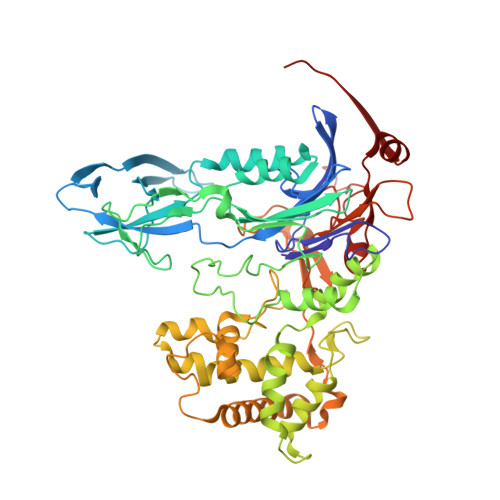
Structural and kinetic studies on ligand binding in wild-type and active-site mutants of penicillin acylase.
Alkema, W.B.L., Hensgens, C.M.H., Snijder, H.J., Keizer, E., Dijkstra, B.W., Janssen, D.B.(2004) Protein Eng Des Sel 17: 473-480
- PubMed: 15254299 Search on PubMed
- DOI: https://doi.org/10.1093/protein/gzh057
- Primary Citation Related Structures:
1JX9, 1K5Q, 1K5S, 1K7D, 1KEC - PubMed Abstract:
Penicillin acylase catalyses the condensation of Calpha-substituted phenylacetic acids with beta-lactam nucleophiles, producing semi-synthetic beta-lactam antibiotics. For efficient synthesis a low affinity for phenylacetic acid and a high affinity for Calpha-substituted phenylacetic acid derivatives is desirable. We made three active site mutants, alphaF146Y, betaF24A and alphaF146Y/betaF24A, which all had a 2- to 10-fold higher affinity for Calpha-substituted compounds than wild-type enzyme. In addition, betaF24A had a 20-fold reduced affinity for phenylacetic acid. The molecular basis of the improved properties was investigated by X-ray crystallography. These studies showed that the higher affinity of alphaF146Y for (R)-alpha-methylphenylacetic acid can be explained by van der Waals interactions between alphaY146:OH and the Calpha-substituent. The betaF24A mutation causes an opening of the phenylacetic acid binding site. Only (R)-alpha-methylphenylacetic acid, but not phenylacetic acid, induces a conformation with the ligand tightly bound, explaining the weak binding of phenylacetic acid. A comparison of the betaF24A structure with other open conformations of penicillin acylase showed that betaF24 has a fixed position, whereas alphaF146 acts as a flexible lid on the binding site and reorients its position to achieve optimal substrate binding.
- Department of Biochemistry, Groningen Biomolecular Sciences and Biotechnology Institute, University of Groningen, Nijenborgh 4, 9747 AG, The Netherlands.
Organizational Affiliation: