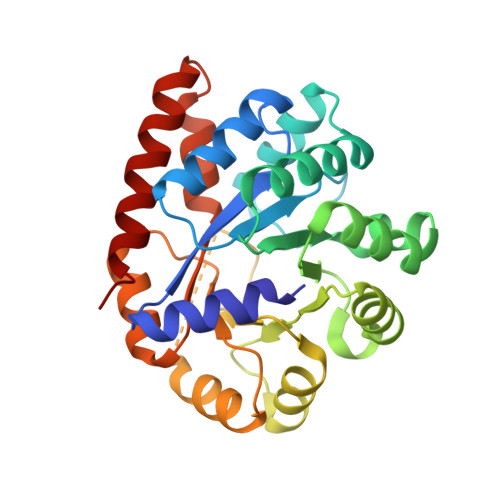
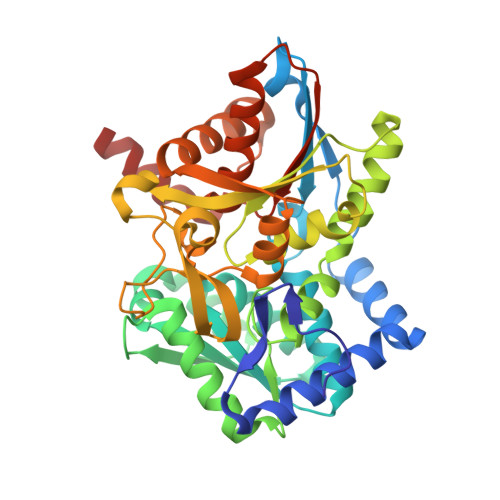
Crystal structures of a new class of allosteric effectors complexed to tryptophan synthase.
Weyand, M., Schlichting, I., Marabotti, A., Mozzarelli, A.(2002) J Biological Chem 277: 10647-10652
- PubMed: 11756456 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M111285200
- Primary Citation Related Structures:
1K3U, 1K7E, 1K7F - PubMed Abstract:
Tryptophan synthase is a bifunctional alpha(2)beta(2) complex catalyzing the last two steps of l-tryptophan biosynthesis. The natural substrates of the alpha-subunit indole- 3-glycerolphosphate and glyceraldehyde-3-phosphate, and the substrate analogs indole-3-propanolphosphate and dl-alpha-glycerol-3-phosphate are allosteric effectors of the beta-subunit activity. It has been shown recently, that the indole-3-acetyl amino acids indole-3-acetylglycine and indole-3-acetyl-l-aspartic acid are both alpha-subunit inhibitors and beta-subunit allosteric effectors, whereas indole-3-acetyl-l-valine is only an alpha-subunit inhibitor (Marabotti, A., Cozzini, P., and Mozzarelli, A. (2000) Biochim. Biophys. Acta 1476, 287-299). The crystal structures of tryptophan synthase complexed with indole-3-acetylglycine and indole-3-acetyl-l-aspartic acid show that both ligands bind to the active site such that the carboxylate moiety is positioned similarly as the phosphate group of the natural substrates. As a consequence, the residues of the alpha-active site that interact with the ligands are the same as observed in the indole 3-glycerolphosphate-enzyme complex. Ligand binding leads to closure of loop alphaL6 of the alpha-subunit, a key structural element of intersubunit communication. This is in keeping with the allosteric role played by these compounds. The structure of the enzyme complex with indole-3-acetyl-l-valine is quite different. Due to the hydrophobic lateral chain, this molecule adopts a new orientation in the alpha-active site. In this case, closure of loop alphaL6 is no longer observed, in agreement with its functioning only as an inhibitor of the alpha-subunit reaction.
- Max-Planck-Institut für Molekulare Physiologie, Abteilung für Physikalische Biochemie, D-44227 Dortmund, Germany.
Organizational Affiliation: