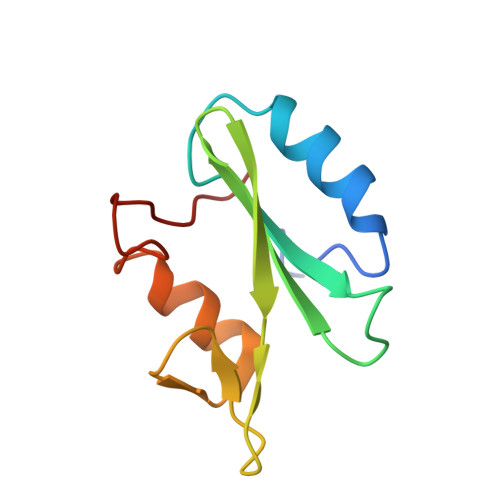
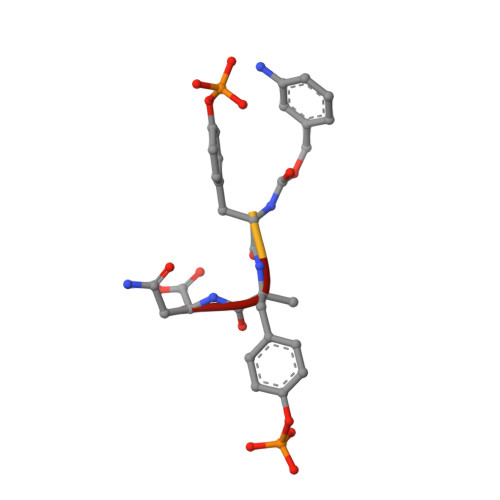
Crystal structures of the SH2 domain of Grb2: highlight on the binding of a new high-affinity inhibitor.
Nioche, P., Liu, W.Q., Broutin, I., Charbonnier, F., Latreille, M.T., Vidal, M., Roques, B., Garbay, C., Ducruix, A.(2002) J Mol Biology 315: 1167-1177
- PubMed: 11827484 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.5299
- Primary Citation Related Structures:
1JYQ, 1JYR, 1JYU - PubMed Abstract:
The activation of growth factor receptors induces phosphorylation of tyrosine residues in its C-terminal part, creating binding sites for SH2 domain-containing proteins. Grb2 is a protein that recruits Sos, the exchange factor for Ras. Recruitment of Sos allows for Ras activation and subsequent signal transmission. This promotes translocation of MAP kinases into the nucleus and activation of early transcription factors. Grb2, a 25 kDa protein, is composed of one SH2 domain surrounded by two SH3 domains. The SH2 domain of Grb2 binds to class II phosphotyrosyl peptides with the consensus sequence pYXNX. Thus, Grb2 is a good example of a bifunctional adaptor protein that brings proteins into close proximity, allowing signal transduction through proteins located in different compartments. To explore the interactions between Grb2 and phosphorylated ligands, we have solved the crystal structure of complexes between the Grb2-SH2 domain and peptides corresponding to Shc-derived sequences. Two structures are described: the Grb2-SH2 domain in complex with PSpYVNVQN at 1.5 A; and the Grb2-SH2 domain in complex with mAZ*-pY-(alphaMe)pY-N-NH2 pseudo-peptide, at 2 A. Both are compared to an unliganded SH2 structure determined at 2.7 A which, interestingly enough, forms a dimer through two swapping subdomains from two symmetry-related molecules. The nanomolar affinity of the mAZ-pY-(alphaMe)pY-N-NH2 pseudo-peptide for Grb2-SH2 is related to new interactions with non- conserved residues. The design of Grb2-SH2 domain inhibitors that prevent interaction with tyrosine kinase proteins or other adaptors like Shc or IRS1 should provide a means to interrupt the Ras signaling pathway. Newly synthesized pseudo-peptides exhibit nanomolar affinities for the Grb2-SH2 domain. It will then be possible to design new inhibitors with similar affinity and simpler chemical structures.
- Laboratoire de Cristallographie et RMN biologiques; UMR 8015 CNRS, Faculté de Pharmacie, Université René Descartes, 4, Avenue de l'Observatorie, 75270 Paris cedex 06, France.
Organizational Affiliation: