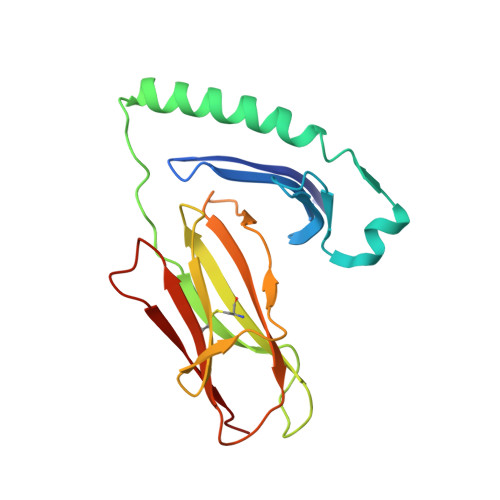
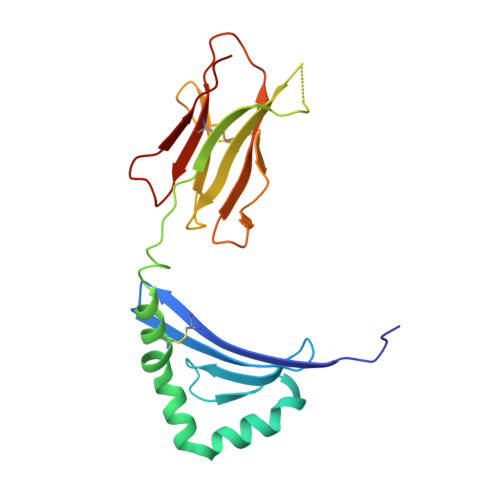
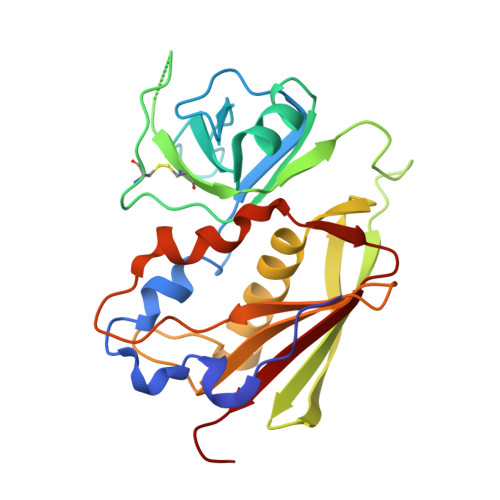
Structural, energetic, and functional analysis of a protein-protein interface at distinct stages of affinity maturation
Sundberg, E.J., Andersen, P.S., Schlievert, P.M., Karjalainen, K., Mariuzza, R.A.(2003) Structure 11: 1151-1161
- PubMed: 12962633 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(03)00187-4
- Primary Citation Related Structures:
1JWM, 1JWS, 1JWU - PubMed Abstract:
Due to a paucity of studies that synthesize structural, energetic, and functional analyses of a series of protein complexes representing distinct stages in an affinity maturation pathway, the biophysical basis for the molecular evolution of protein-protein interactions is poorly understood. Here, we combine crystal structures and binding-free energies of a series of variant superantigen (SAG)-major histocompatibility complex (MHC) class II complexes exhibiting increasingly higher affinity to reveal that this affinity maturation pathway is controlled largely by two biophysical factors: shape complementarity and buried hydrophobic surface. These factors, however, do not contribute equivalently to the affinity maturation of the interface, as the former dominates the early steps of the maturation process while the latter is responsible for improved binding in later steps. Functional assays reveal how affinity maturation of the SAG-MHC interface corresponds to T cell activation by SAGs.
- Center for Advanced Research in Biotechnology, W.M. Keck Laboratory for Structural Biology, University of Maryland Biotechnology Institute, 9600 Gudelsky Drive, Rockville, MD 20850, USA. sundberg@umbi.umd.edu
Organizational Affiliation: