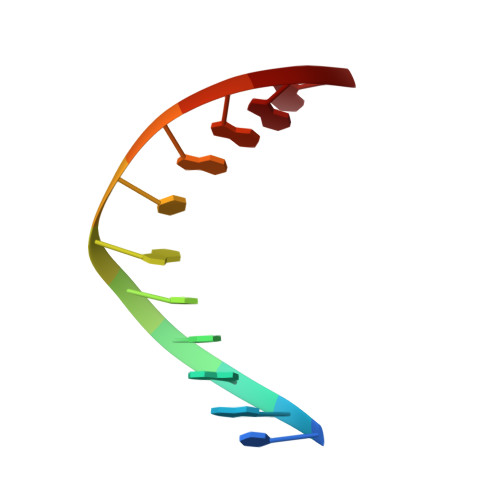
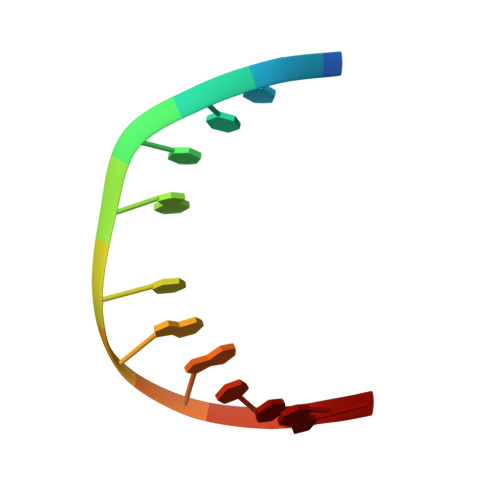
Solution structure of a trans-opened (10S)-dA adduct of (+)-(7S,8R,9S,10R)-7,8-dihydroxy-9,10-epoxy-7,8,9,10-tetrahydrobenzo[a]pyrene in a fully complementary DNA duplex: evidence for a major syn conformation.
Pradhan, P., Tirumala, S., Liu, X., Sayer, J.M., Jerina, D.M., Yeh, H.J.(2001) Biochemistry 40: 5870-5881
- PubMed: 11352722 Search on PubMed
- DOI: https://doi.org/10.1021/bi002896q
- Primary Citation Related Structures:
1JDG - PubMed Abstract:
Two-dimensional NMR was used to determine the solution structure of an undecanucleotide duplex, d(CGGTCACGAGG).d(CCTCGTGACCG), in which (+)-(7S,8R,9S,10R)-7,8-dihydroxy-9,10-epoxy-7,8,9,10-tetrahydrobenzo[a]pyrene is covalently bonded to the exocyclic N(6)() amino group of the central deoxyadenosine, dA(6), through trans addition at C10 of the epoxide (to give a 10S adduct). The present study represents the first NMR structure of a benzo[a]pyrene (10S)-dA adduct in DNA with a complementary T opposite the modified dA. Exchangeable and nonexchangeable protons of the modified duplex were assigned by the use of TOCSY (in D(2)O) and NOESY spectra (in H(2)O and D(2)O). Sequential NOEs expected for a B-type DNA conformation with typical Watson-Crick base pairing are observed along the duplex, except at the lesion site. We observed a strong intraresidue NOE cross-peak between H1' and H8 of the modified dA(6). The sugar H2' and H2' ' of dC(5) lacked NOE cross-peaks with H8 of dA(6) but showed weak interactions with H2 of dA(6) instead. In addition, the chemical shift of the H8 proton (7.51 ppm) of dA(6) appears at a higher field than that of H2 (8.48 ppm). These NOE and chemical shift data for the dA(6) base protons are typical of a syn glycosidic bond at the modified base. Restrained molecular dynamics/energy minimization calculations show that the hydrocarbon is intercalated from the major groove on the 3'-side of the modified base between base pairs A(6)-T(17) and C(7)-G(16) and confirm the syn glycosidic angle (58 degrees ) of the modified dA(6). In the syn structure, a weak A-T hydrogen bond is possible between the N3-H proton of T(17) and N7 of dA(6) (at a distance of 3.11 A), whereas N1, the usual hydrogen bonding partner for N3-H of T when dA is in the anti conformation, is 6.31 A away from this proton. The 10(S)-dA modified DNA duplex remains in a right-handed helix, which bends in the direction of the aliphatic ring of BaP at about 42 degrees from the helical axis. ROESY experiments provided evidence for interconversion between the major, syn conformer and a minor, possibly anti, conformer.
- Laboratory of Bioorganic Chemistry, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, Maryland 20892, USA.
Organizational Affiliation: