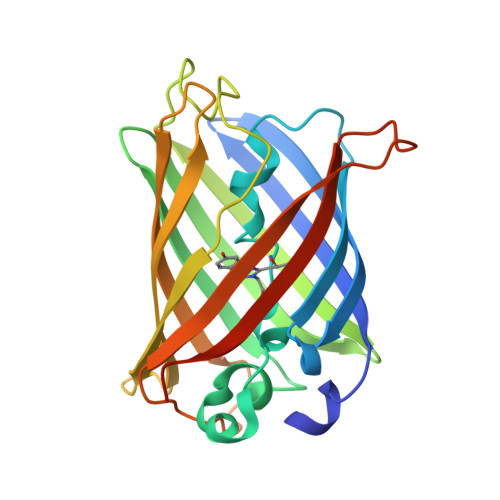
Green Fluorescent Protein Variants as Ratiometric Dual Emission pH Sensors. 1. Structural Characterization and Preliminary Application.
Hanson, G.T., McAnaney, T.B., Park, E.S., Rendell, M.E.P., Yarbrough, D.K., Chu, S., Xi, L., Boxer, S.G., Montrose, M.H., Remington, S.J.(2002) Biochemistry 41: 15477-15488
- PubMed: 12501176 Search on PubMed
- DOI: https://doi.org/10.1021/bi026609p
- Primary Citation Related Structures:
1JBY, 1JBZ - PubMed Abstract:
Novel dual emission, pH-sensitive variants of the green fluorescent protein (GFP) have been constructed and are suitable for ratiometric emission measurements in vivo. This new class of GFPs, termed deGPFs, results from substitution of wild-type residue 65 with threonine and residues 148 and/or 203 with cysteine. deGFPs display pK(a) values ranging from 6.8 to 8.0 and emission that switches from a green form (lambda(max) approximately 515 nm) to a blue form (lambda(max) approximately 460 nm) with acidifying pH. In this report we analyze in most detail the deGFP1 variant (S65T/H148G/T203C, pK(a) approximately 8.0) and the deGFP4 variant (S65T/C48S/H148C/T203C, pK(a) approximately 7.3). In the following paper [McAnaney, T. B., Park, E. S., Hanson, G. T., Remington, S. J., and Boxer, S. G. (2002) Biochemistry 41, 15489-15494], data obtained by ultrafast fluorescence upconversion spectroscopy can be described by a kinetic model that includes an excited-state proton-transfer pathway at high pH but not at low pH. Crystal structure analyses of deGFP1 at high-pH and low-pH conformations were performed to elucidate the basis for the dual emission characteristics. At low pH the structure does not contain a hydrogen bond network that would support rapid transfer of a proton from the excited state of the neutral chromophore to a suitable acceptor; hence blue emission is observed. At high pH, backbone rearrangements induced by changes in the associated hydrogen bond network permit excited-state proton transfer from the excited state of the neutral chromophore to the bulk solvent via Ser147 and bound water molecules, resulting in green emission from the anionic chromophore. Comparative analysis suggests that the basis for dual emission is elimination of the wild-type proton-transfer network by the S65T substitution, a general reduction in hydrogen-bonding opportunities, and a concomitant increase in the hydrophobic nature of the chromophore environment resulting from the cysteine substitutions. We evaluated the suitability of the deGFP4 variant for intracellular pH measurements in mammalian cells by transient expression in PS120 fibroblasts. The responses of deGFP4 and a commercially available pH-sensitive dye, SNARF-1, to changes in pH were compared in the same cells. Results show that the dynamic range of the emission ratio change is comparable between the two pH sensors over the range examined. Two-photon excitation was found to elicit a better deGFP4 fluorescent signal above cellular autofluorescence when compared to conventional confocal microscopy. Given their favorable optical characteristics, suitable pK(a)'s for the physiological pH range, and suitability for ratiometric measurements, dual emission GFPs should make excellent probes for studying pH in vivo.
- Department of Chemistry, Institute of Molecular Biology, University of Oregon, Eugene, OR 97403-1229, USA.
Organizational Affiliation: