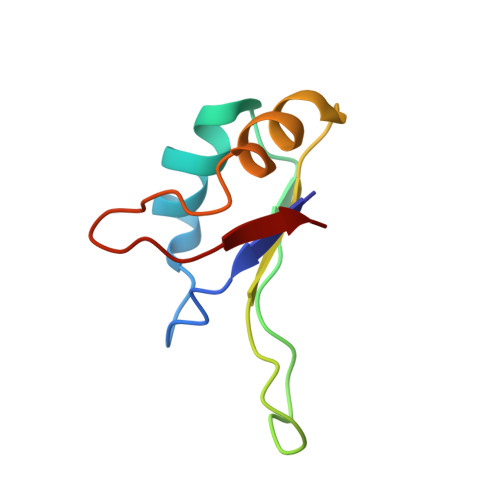
Structure of the C-terminal RNA-binding domain of hnRNP D0 (AUF1), its interactions with RNA and DNA, and change in backbone dynamics upon complex formation with DNA.
Katahira, M., Miyanoiri, Y., Enokizono, Y., Matsuda, G., Nagata, T., Ishikawa, F., Uesugi, S.(2001) J Mol Biology 311: 973-988
- PubMed: 11531333
- DOI: https://doi.org/10.1006/jmbi.2001.4862
- Primary Citation of Related Structures:
1IQT - PubMed Abstract:
Heterogeneous nuclear ribonucleoprotein (hnRNP) D0 has two ribonucleoprotein (RNP) -type RNA-binding domains (RBDs), each of which can specifically bind to the UUAG-sequence. hnRNP D0 also binds specifically to single-stranded d(TTAGGG)(n), the human telomeric DNA repeat. We have already reported the structure and interactions with RNA of the N-terminal RBD (RBD1). Here, the structure of the C-terminal RBD (RBD2) determined by NMR is presented. It folds into a compact alpha beta structure comprising an antiparallel beta-sheet packed against two alpha-helices, which is characteristic of RNP-type RBDs. In addition to the four beta-strands commonly found in RNP-type RBDs, an extra beta-strand, termed beta 4(-), was found just before the fourth beta-strand, yielding a five-stranded beta-sheet. Candidate residues of RBD2 involved in the interactions with RNA were identified by chemical shift perturbation analysis. Perturbation was detected on the beta-sheet side, not on the opposite alpha-helix side, as observed for RBD1. It is notable that the beta 4(-) to beta 4 region of RBD2 is involved in the interactions in contrast to the case of RBD1. The chemical shift perturbation analysis also showed that RBD2 interacts with DNA in essentially the same way as with RNA. Changes in the backbone dynamics upon complex formation with DNA were examined by means of model free analysis of relaxation data. In free RBD2, the beta 4(-) to beta 4 region exhibits slow conformational exchange on the milli- to microsecond time scale. The exchange is quenched upon complex formation. The flexibility of free RBD2 may be utilized in the recognition process by allowing different conformational states to be accessed and facilitating induced fit. Additionally, faster flexibility on the nano- to picosecond time scale was observed for loop 3 located between beta 2 and beta 3 in free RBD2, which is retained by the complex as well.
- Department of Environment and Natural Sciences, Graduate School of Environment and Information Sciences, 79-7 Tokiwadai, Hodogaya-ku, Yokohama, 240-8501, Japan
Organizational Affiliation: