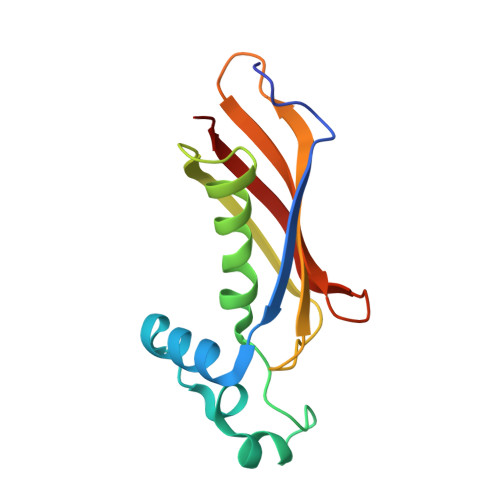
Crystal structure of the (R)-specific enoyl-CoA hydratase from Aeromonas caviae involved in polyhydroxyalkanoate biosynthesis
Hisano, T., Tsuge, T., Fukui, T., Iwata, T., Miki, K., Doi, Y.(2003) J Biological Chem 278: 617-624
- PubMed: 12409309 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M205484200
- Primary Citation Related Structures:
1IQ6 - PubMed Abstract:
The (R)-specific enoyl coenzyme A hydratase ((R)-hydratase) from Aeromonas caviae catalyzes the addition of a water molecule to trans-2-enoyl coenzyme A (CoA), with a chain-length of 4-6 carbons, to produce the corresponding (R)-3-hydroxyacyl-CoA. It forms a dimer of identical subunits with a molecular weight of about 14,000 and is involved in polyhydroxyalkanoate (PHA) biosynthesis. The crystal structure of the enzyme has been determined at 1.5-A resolution. The structure of the monomer consists of a five-stranded antiparallel beta-sheet and a central alpha-helix, folded into a so-called "hot dog" fold, with an overhanging segment. This overhang contains the conserved residues including the hydratase 2 motif residues. In dimeric form, two beta-sheets are associated to form an extended 10-stranded beta-sheet, and the overhangs obscure the putative active sites at the subunit interface. The active site is located deep within the substrate-binding tunnel, where Asp(31) and His(36) form a catalytic dyad. These residues are catalytically important as confirmed by site-directed mutagenesis and are possibly responsible for the activation of a water molecule and the protonation of a substrate molecule, respectively. Residues such as Leu(65) and Val(130) are situated at the bottom of the substrate-binding tunnel, defining the preference of the enzyme for the chain length of the substrate. These results provide target residues for protein engineering, which will enhance the significance of this enzyme in the production of novel PHA polymers. In addition, this study provides the first structural information of the (R)-hydratase family and may facilitate further functional studies for members of the family.
- RIKEN Institute, 2-1 Hirosawa, Wako, Saitama 351-0198, Japan. hisano@postman.riken.go.jp
Organizational Affiliation: