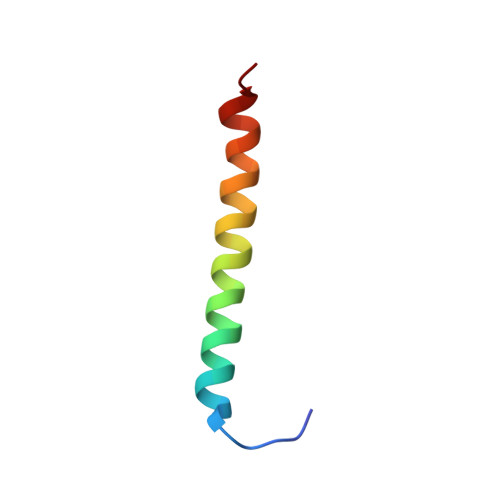
Solution NMR structure and folding dynamics of the N terminus of a rat non-muscle alpha-tropomyosin in an engineered chimeric protein.
Greenfield, N.J., Huang, Y.J., Palm, T., Swapna, G.V., Monleon, D., Montelione, G.T., Hitchcock-DeGregori, S.E.(2001) J Mol Biology 312: 833-847
- PubMed: 11575936 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.4982
- Primary Citation Related Structures:
1IHQ - PubMed Abstract:
Tropomyosin is an alpha-helical coiled-coil protein that aligns head-to-tail along the length of the actin filament and regulates its function. The solution structure of the functionally important N terminus of a short 247-residue non-muscle tropomyosin was determined in an engineered chimeric protein, GlyTM1bZip, consisting of the first 19 residues of rat short alpha-tropomyosin and the last 18 residues of the GCN4 leucine zipper. A gene encoding GlyTM1bZip was synthesized, cloned and expressed in Escherichia coli. Triple resonance NMR spectra were analyzed with the program AutoAssign to assign its backbone resonances. Multidimensional nuclear Overhauser effect spectra, X-filtered spectra and (3)J(H(N)-H(alpha)) scalar coupling were analyzed using AutoStructure. This is the first application of this new program to determine the three-dimensional structure of a symmetric homodimer and a structure not previously reported. Residues 7-35 in GlyTM1bZip form a coiled coil, but neither end is helical. Heteronuclear (15)N-(1)H nuclear Overhauser effect data showed that the non-helical N-terminal residues are flexible. The (13)C' chemical shifts of the coiled-coil backbone carbonyl groups in GlyTM1bZip showed a previously unreported periodicity, where resonances arising from residues at the coiled-coil interface in a and d positions of the heptad repeat were displaced relatively upfield and those arising from residues in c positions were displaced relatively downfield. Heteronuclear single quantum coherence spectra, collected as a function of temperature, showed that cross-peaks arising from the alpha-helical backbone and side-chains at the coiled-coil interface broadened or shifted with T(M) values approximately 20 degrees C lower than the loss of alpha-helix measured by circular dichroism, suggesting the presence of a folding intermediate. The side-chain of Ile14, a residue essential for binding interactions, exhibited multiple conformations. The conformational flexibility of the N termini of short tropomyosins may be important for their binding specificity.
- Department of Neuroscience and Cell Biology, UMDNJ-Robert Wood Johnson Medical School, Piscataway, NJ 08854-5635, USA. greenfie@rwja.umdnj.edu
Organizational Affiliation: