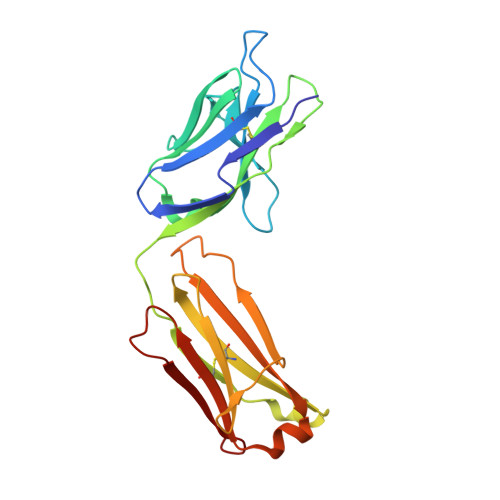
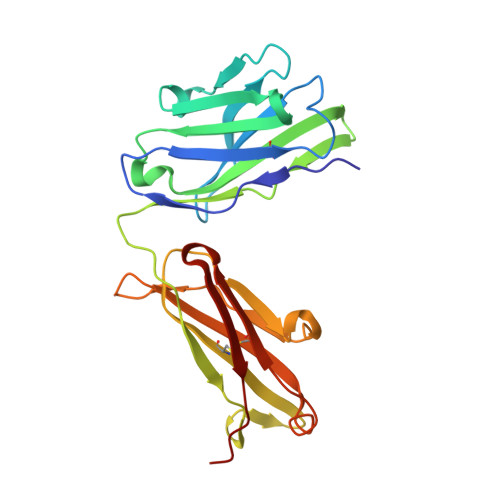
Crystal structure of a cocaine-binding antibody.
Larsen, N.A., Zhou, B., Heine, A., Wirsching, P., Janda, K.D., Wilson, I.A.(2001) J Mol Biology 311: 9-15
- PubMed: 11469854 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.4839
- Primary Citation Related Structures:
1I7Z - PubMed Abstract:
Murine monoclonal antibody GNC92H2 was elicited by active immunization with a cocaine immunoconjugate and binds free cocaine with excellent specificity and moderate affinity. Improvement of affinity, as well as humanization of GNC92H2, would be advantageous in immunopharmacotherapy for cocaine addiction, and for emergency cases of drug overdose. Toward this end, the crystal structure of an engineered murine-human chimeric Fab of GNC92H2 complexed with cocaine was determined at 2.3 A resolution. Structural analysis reveals a binding pocket with high shape and charge complementarity to the cocaine framework, which explains the specificity for cocaine, as opposed to the pharmacologically inactive cocaine metabolites. Importantly, the structure provides a foundation for mutagenesis to enhance the binding affinity for cocaine and potent cocaine derivatives, such as cocaethylene, and for additional humanization of the antibody.
- Department of Molecular Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: