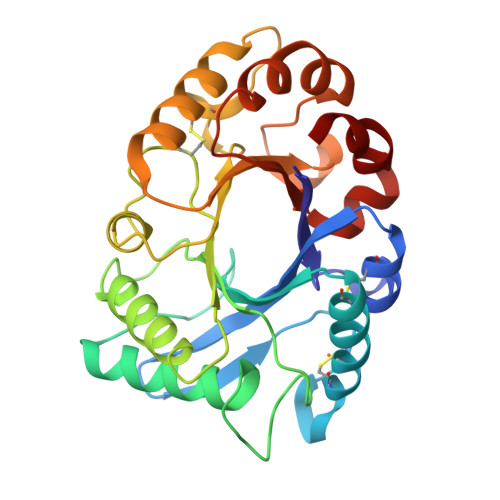
Crystal structures of hevamine, a plant defence protein with chitinase and lysozyme activity, and its complex with an inhibitor.
Terwisscha van Scheltinga, A.C., Kalk, K.H., Beintema, J.J., Dijkstra, B.W.(1994) Structure 2: 1181-1189
- PubMed: 7704528 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(94)00120-0
- Primary Citation Related Structures:
1HVQ - PubMed Abstract:
Hevamine is a member of one of several families of plant chitinases and lysozymes that are important for plant defence against pathogenic bacteria and fungi. The enzyme can hydrolyze the linear polysaccharide chains of chitin and peptidoglycan. A full understanding of the structure/function relationships of chitinases might facilitate the production of transgenic plants with increased resistance towards a wide range of pathogens. The crystal structure of hevamine has been determined to a resolution of 2.2 A, and refined to an R-factor of 0.169. The enzyme possesses a (beta alpha)8-barrel fold. An inhibitor binding study shows that the substrate-binding cleft is located at the carboxy-terminal end of the beta-barrel, near the conserved Glu127. Glu127 is in a position to act as the catalytic proton donor, but no residue that might stabilize a positively charged oxocarbonium ion intermediate was found. A likely mechanism of substrate hydrolysis is by direct attack of a water molecule on the C1 atom of the scissile bond, resulting in inversion of the configuration at C1. The structure of hevamine shows a completely new lysozyme/chitinase fold and represents a new class of polysaccharide-hydrolyzing (beta alpha)8-barrel enzymes. Because the residues conserved in the family to which hevamine belongs are important for maintaining the structure of the (beta alpha)8-barrel, all members of the family, including fungal, bacterial and insect chitinases, are likely to share this architecture. The crystal structure obtained provides a basis for protein engineering studies in this family of chitinases.
- BIOSON Research Institute, University of Groningen, The Netherlands.
Organizational Affiliation: